Outline
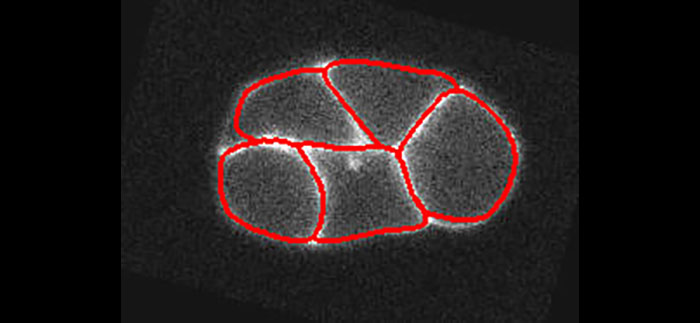
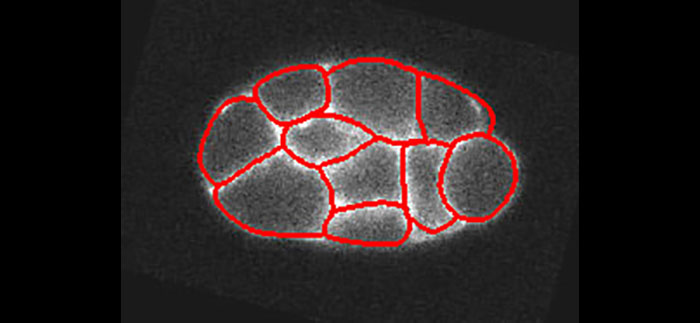
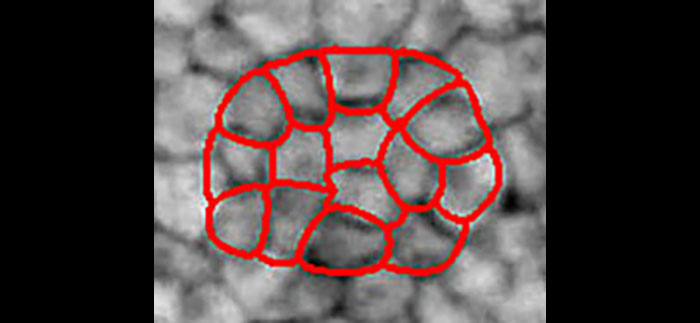
We present a new active contour to segment cell aggregates. We describe it by a smooth tessellation that is attracted toward the cell membranes. Our approach relies on subdivision schemes that are tightly linked to the theory of wavelets. The shape is encoded by control points grouped in tiles. The smooth and continuously defined boundary of each tile is generated by recursively applying a refinement process to its control points. We deform the smooth tessellation in a global manner using a ridge-based energy that we have designed for that purpose. By construction, cells are segmented without overlap and the tessellation structure is maintained even on dim membranes. Leakage, which afflicts usual image-processing methods (e.g., watershed), is thus prevented. We validate our framework on both synthetic and real microscopy images, showing that the proposed method is robust to membrane gaps and to high levels of noise.
Reference
A. Badoual, A. Galan, D. Sage, and M. Unser, "Deforming tessellations for the segmentation of cell aggregates," Proceedings of the Sixteenth IEEE International Symposium on Biomedical Imaging: From Nano to Macro (ISBI'19), Venice, Italian Republic, April 8-11, 2019, pp. 1013-1017.
Open Source Plugin: AC_ActiveTessellations
The method is implemented as a plugin for the Icy bioimaging platform.
Distribution
Demo
Test image and settings
Condition of use
The software is freely available for research purposes. However, it should not be redistributed without the consent of the authors. We expect the user to include a citation of this publication whenever presenting or publishing results that are based on the Icy plugin AC_ActiveTessellations.