The double helix PSF: implementation and localization applications
Spring 2013
Master Semester Project
Project: 00253
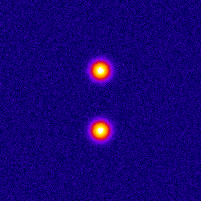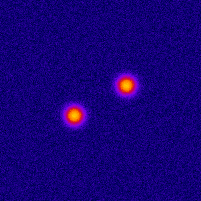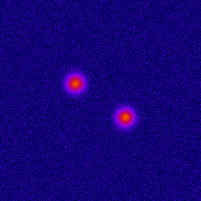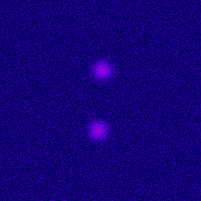
Photo-activated Localization Microscopy (PALM) provides an attractive framework for studying the living cell at a nanometer scale. It is based on exciting a sparse set of fluorophores at each time frame, and on fitting the image of each fluorophore with a point-spread function (PSF). In order to improve the axial localization accuracy of this method, one can design a PSF that exhibits larger sensitivity to the depth value. One such design is the double helix PSF, which consists of two lobes that rotate as a function of the point-source depth value.
The purpose of this project is to investigate the properties of the double-helix PSF within the context of localization microscopy. The main stages of the project are: numerical optimization for the phase function that leads to the double helix pattern, implementation of the PSF model in Java, and 3-D localization analysis for different acquisition conditions: sub-pixel positions, depth values, refractive index mismatch, and noise.
Background in optics and programming is recommended.
- Supervisors
- Hagai Kirshner, hagai.kirshner@epfl.ch, 31136, BM 4.142
- Michael Unser, michael.unser@epfl.ch, 021 693 51 75, BM 4.136