| CONTENTS |
| Zsuzsanna Püspöki | Project |
| Life Science Summer Research Program, EPFL | August 2009 |
Cells are mostly transparent in their natural state and the molecules constituting them are optically indistinguishable from one another. In order to identify the proteins of interest, biologists try to attach bright markers to them. Those markers indicate the position of the proteins while having a very low impact on their functions. Fluorescent proteins are good candidates to be such a marker. Our project concerns the detection of fluorescence or bright particles having different sizes in biological images. The particles can be modeled as being circular, and our goal is to detect their centers and radii even if they are elongated or damaged by noise.
Mexican-hat detectors can be realized based on the fact that radially-uniform box splines with uniform scale vectors resemble isotropic Gaussians. In order to specify such a detector, we consider the difference of two isotropic radially-uniform box splines. The box spline detector and the corresponding detector profile along a given direction is shown in Fig. 1. While the detector suppress uniform-intensity regions, a large response is generated by the discontinuities along the radial direction. A typical detector characteristic is shown in Fig. 2.
The fundamental aspect of our algorithm is its space-variance which was used to continuously adjust the size of the detector during the optimization procedure (golden section search). We note that the ideas used in the above cell detection algorithm can also be transposed to similar image analysis applications. As a result of my work an ImageJ plugin was created which can be used for particle detection.
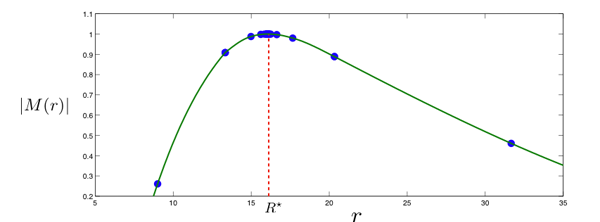
Fig. 1
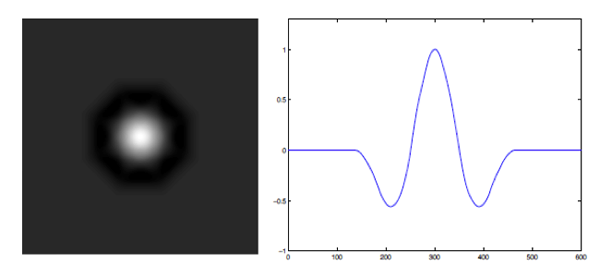
Fig. 2
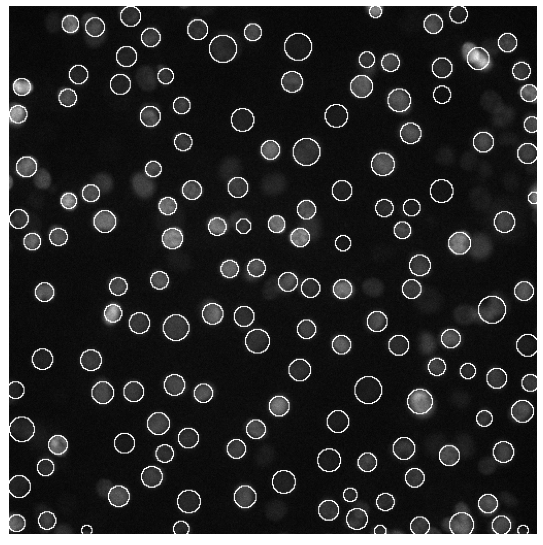

Fig. 3
© 2022 EPFL • webmaster.big@epfl.ch • 11.08.2022