| EPFL > BIG > Teaching > Student Projects > Completed Projects > Daniel Schmitter |
| CONTENTS |
|
Student Projects |
Daniel Schmitter | Semester Master Project |
Life Science, EPFL | June 2011 |
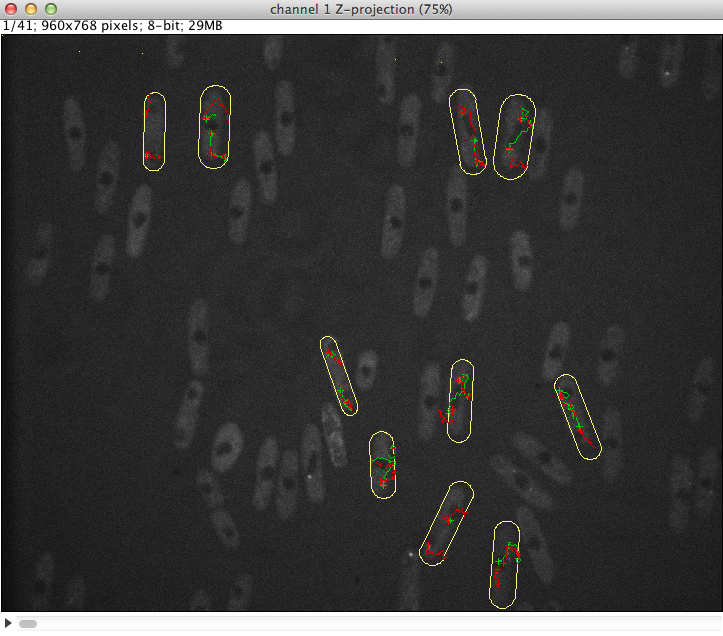
The goal of this project was to develop a model to quantify the asymmetry of spindle pole bodies in yeast cells (S. Pombe) and to implement it as an ImageJ plugin. Plugins for dynamic (time lapse sequences) as well as for static (single) images were developed. The images can be either 2D or 3D. The processing steps leading to quantitaion are the follwing:
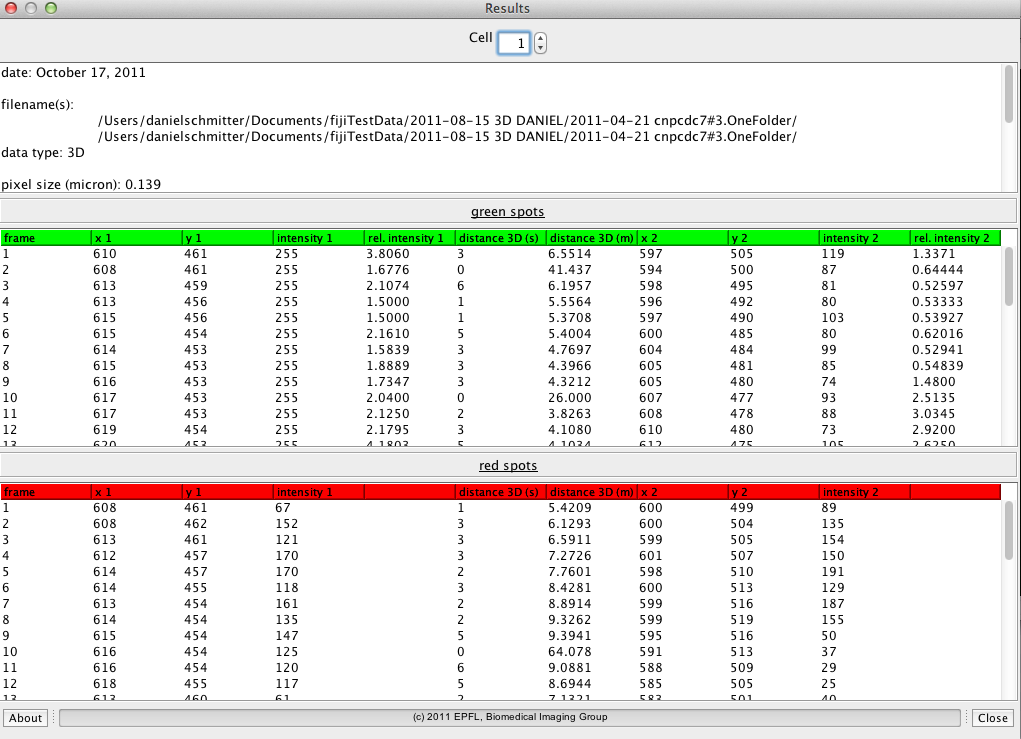
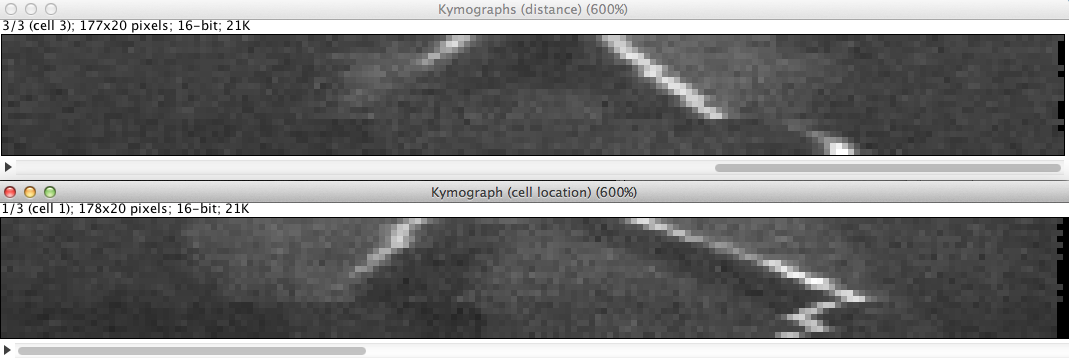
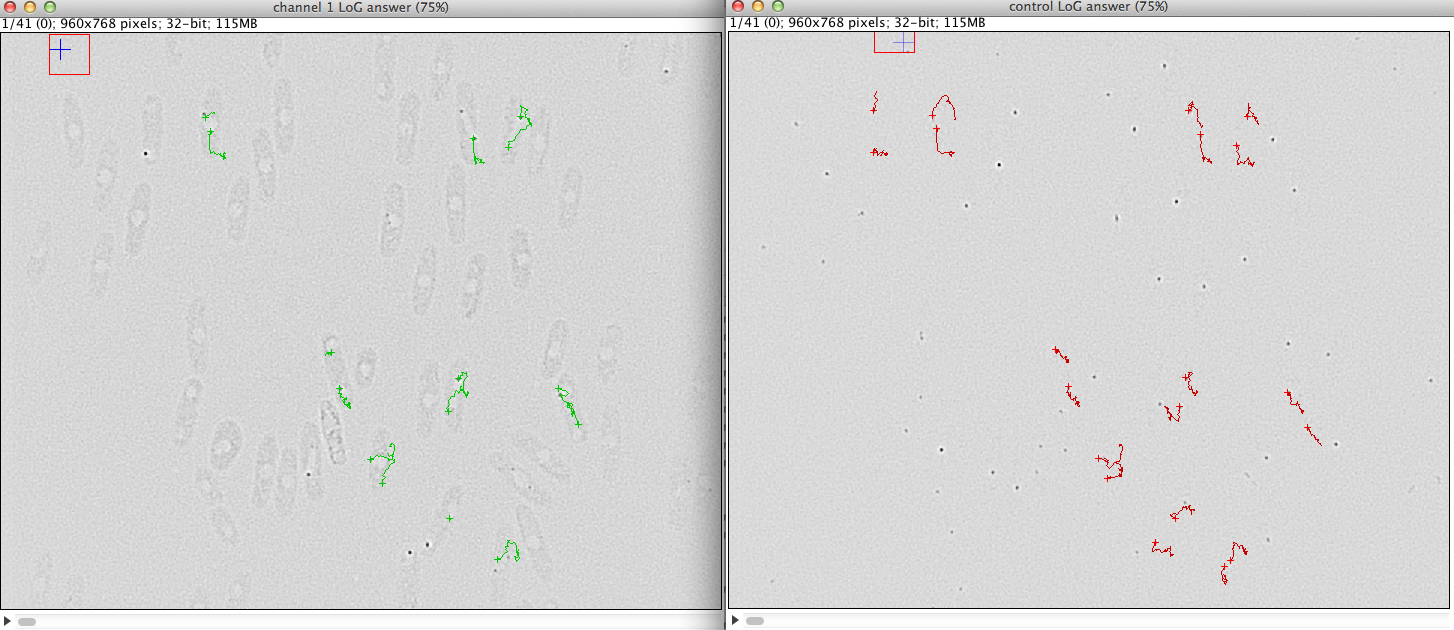
Up to 4 proteins (2 proteins of interest and 2 reference proteins) can be tracked per cell and the number of cells to segment is unlimited. The results are dis- played in a table and 2 di erent kymographs can be generated. The results can be saved in an tab-delimited text le for further analysis with other software (excel, matlab, etc.). After denoising and LoG- ltering of the raw data, the proteins are detected. The tracking is done through a dynamic programing algorithm which allows to nd the trajectory even through very noisy sequences. For the cell segmenta- tion an active contour model was developed which adapts to the rod shaped cell structure. A plugin just to segment rod shaped cells was individually developed.
© 2022 EPFL • webmaster.big@epfl.ch • 11.08.2022