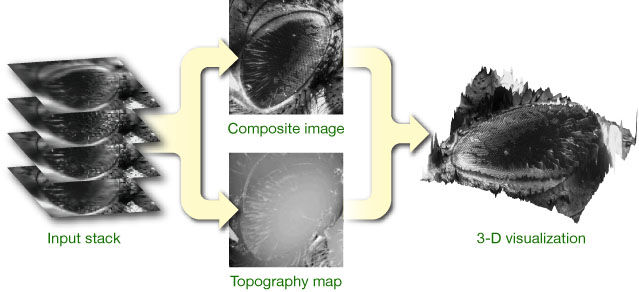
Due to the limited depth of field of brightfield microscopes, it is usually impossible to image large 3-D organisms and objects entirely in focus. By optically sectioning the specimen, however, the in-focus information at the specimen's surface can be acquired over a range of images, which can then be processed to generate a single in-focus image. We have developed different algorithms to achieve this, and provide free software that implements and demonstrates these methods.
The idea is to merge a stack of micrographs taken at different focal positions (aligned along the optical axis) into a single, entirely focused composite image, as illustrated on the right. Traditional extended depth of field algorithms rely on a high-pass criterion that is applied to each image in the stack; in-focus regions are then identifed based on local energy. An efficient approach consists in using a wavelet transform, where a selection rule based on maximum absolute coefficient values leads to good results. Our complex wavelet-based method expands on this concept and demonstrates state-of-the-art performance for this class of algorithms. Unfortunately, the topography provided by these techniques is limited to a map of selected in-focus pixel positions and is inherently discretized along the axial direction, which motivated the development of a more advanced approach based on an image formation model.
Interpretation of the composite extended depth of field specimen can be difficult; to this end we provide software for the 3-D visualization of in-focus image/topography pairs. Such pairs can also be visualized using anaglyphs.
Reference
B. Forster, D. Van De Ville, J. Berent, D. Sage, M. Unser, Extended Depth-of-Focus for Multi-Channel Microscopy Images: A Complex Wavelet Approach Proceedings of the Second IEEE International Symposium on Biomedical Imaging, Arlington VA, USA, 2004.
The complex wavelet transform leads to improved extended depth of field results over other wavelet-based approaches. In order to further improve the quality of the results, we introduced post-processing steps that enforce local smoothness of the topography, avoid saturation and accumulation of noise further improve the quality of the results. In order to process color data efficiently, an RGB to grayscale conversion method that preserves contrast and intensity has been implemented; in this way, the selection of in-focus pixels is performed in a single channel. The color values of the composite image are then retrieved from the original stack.
B. Forster, D. Van De Ville, J. Berent, D. Sage, M. Unser, "Complex Wavelets for Extended Depth-of-Field: A New Method for the Fusion of Multichannel Microscopy Images," Microsc. Res. Tech., 65, September 2004.
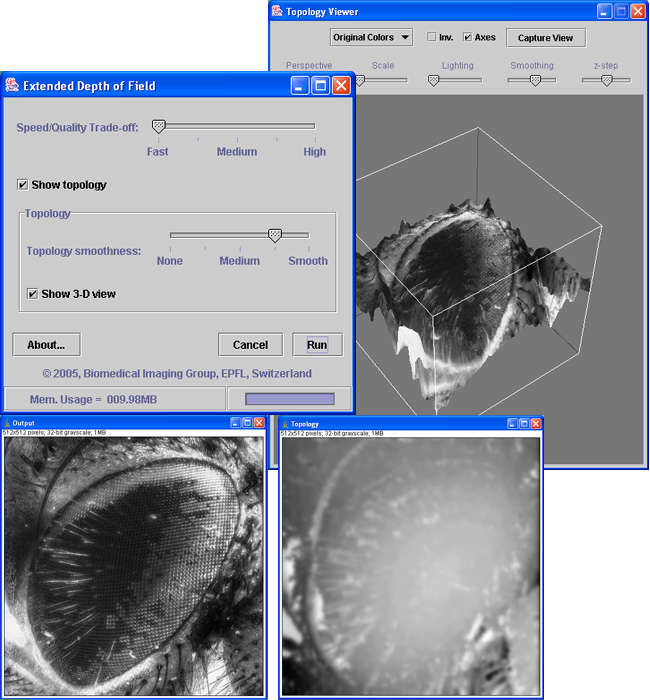
This software implements different wavelet-based algorithms for extended depth of field. Two interfaces are provided for adjusting the algorithm parameters: a simplified, easy-to-use mode and an expert mode that gives access to the complete set of parameters. A 3D topology viewer plugin is inlcuded.
This software was developed by Alex Prudencio, Jesse Berent and Daniel Sage (contact). The 3-D view was developped by Kai Uwe Barthel.
Install and Run itThe plugin is callable from an ImageJ macro, in "easy" mode. The parameters are described and an example is given in EDF-demo.txt
In order to recover an accurate, continuous representation of the specimen topography, extended depth of field is stated as an optimization problem where the texture and topography are jointly estimated in an iterative process. Image formation is modeled as the convolution of a thick specimen model with the microscope's point spread function. The method acts as a deconvolution when the in-focus PSF has a blurring effect, or when the true in-focus position falls in between two optical sections. The resulting in-focus composite image and topography are of very high quality, albeit at a larger computational cost when compared to wavelet-based approaches.
F. Aguet, D. Van De Ville, and M. Unser "Model-based 2.5-D deconvolution for extended depth-of-field in brightfield microscopy," IEEE Trans. Image Process., 17(7), pp. 1144-1153, July 2008.
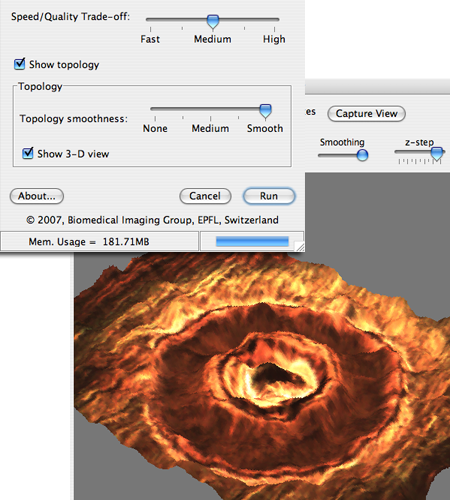
This software implements our new, model-based algorithm for extended depth of field. The following functionalities are provided:
Developed by François Aguet.
MattheddleiteThe images are courtesy of Rick Turner, British Micromount Society. These images was obtained using an Olympus DP12 camera attached to a Wild M7A stereomicroscope. FoV is approx 1mm. The light blue is the mattheddleite, the dark blue linarite, and the white mostly leadhillite. The specimen comes from Red Gill Mine, in the English Lake District. The whole stack is composed of 10 color images of 1024x768 pixels. | |
Image stack of a fly's eyeThe images are courtesy of Philippe Thévenaz, EPFL, Lausanne The whole stack is composed of 32 14-bits images of 1280x1024 pixels. The images have been realigned first using: StackReg our registration software. The huge dataset of original images can be provided on request. |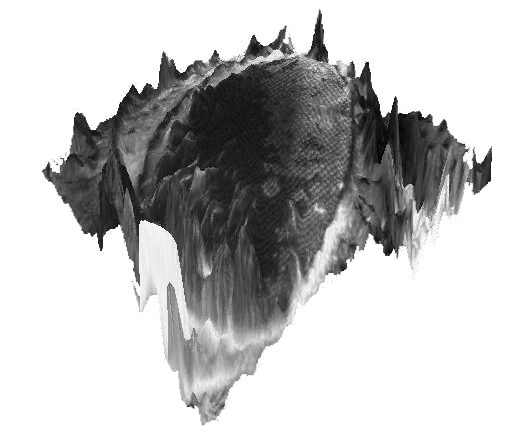  |
Study of the imapact of laser weldingThe images are courtesy of Thomas Sidler, EPFL, Lausanne. The whole stack is composed of 13 color images of 1024x768 pixels. The huge dataset of original images can be provided on request. |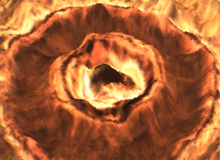  |
Study of the retinal pigment epitheliumThe images are courtesy of Peter Lundh von Leithner and Heba Ahmad, Institute of Ophthalmology, London. The whole stack is composed of 4 color images of 1024x768 pixels. | |
Mouse intestin's Peyer's patchesThe images are courtesy of Nathalie Garin, ISREC, Lausanne. The whole stack is composed of 20 color images of 1996x1450 pixels. Images shows mouse intestin's Peyer's patches. The huge dataset of original images can be provided on request. |  |
PancreasThe images are courtesy of Claude Bonnard, ISREC, Lausanne Stack of images of a pancreas. The huge dataset of original images can be provided on request. |
The Extended Depth of Field plugin implements some other algorithms also mentioned in the reference. The table below shows some performance results comparing the different algorithms.
| Memory | Elapsed Time [s] | ||
|---|---|---|---|
low resolution (256 x 256 pixels, # Slices = 15) |
high resolution (1024x1024 pixels,
#Slices=15) |
||
Sobel |
~3 times the stack size | 3 |
34 |
Variance |
~3 times the stack size | 4 |
56 |
Real Wavelets |
~3 times the stack size | 5 |
83 |
Complex Wavelets |
~8 times the stack size | 23 |
385 |
 |
 |
Another option for 3-D visualization of the topography is to use the Anaglyph ImageJ plugin written by Gabriel Landini. This plugin creates (color or greyscale) red-cyan or red-green anaglyphs, stereo pairs (crossed view) and depth map images from the topography and in-focuse image generated by the Extended Depth of Field plugin. Details and lastest version of the plugin and can be obtained at G. Landini's Software page.