| Sébastien Romain | Semester project |
| Erasmus, EPFL | June 2007 |
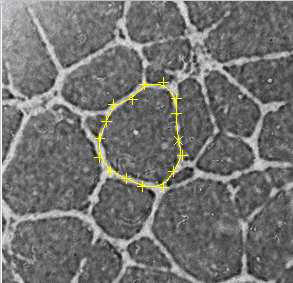
The objective of this project is to develop a new technique for characterizing cells in a certain class of biological images. This enables them to compute parameters such as the cell size, area, velocity and also understand cell growth by comparing across images. This is best achieved by an algorithm that matches a deformable contour to the edges of the objects.
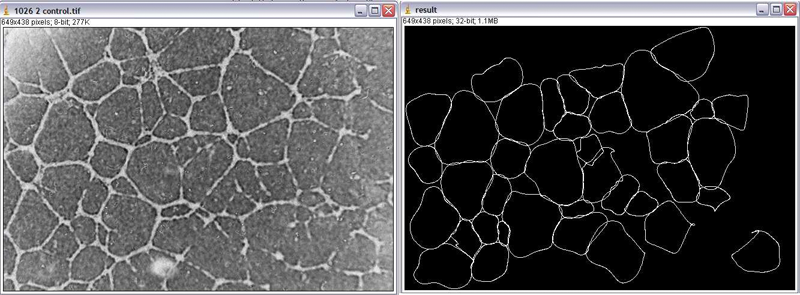
During this project, several methods has been developed and compared to build an algorithm able to match the edge of a selected cell in a grey-scale image. Among all the solutions considered, the edge distance map based snake algorithm is the one that presents the more advantages. At first, it is relatively simple. It use only two energy terms, and can even be run using only the Edge distance-based Energy term. The second strength of this algorithm is that it can work with cell which does not have a contour completely well defined. Some good matching can be made even on cells which had a missing part of the contour.
But this algorithm has also some weakness. First, to build the edge distance map, a good edge information is needed. It requires the use of a steerable filter that is time consuming. The edge distance map requires also a non-negligible time to be built. This time can be reduced by reducing the area of processing to a small area around the cell to match. The second point that can be improved is the fact that the initialization requires the intervention of the user, so at this point, it is still impossible to build an automatic cell matching algorithm with this method. But the ImageJ plugin developed in this project can be an useful tool for a biologist in the sense that it permits to match cells relatively well.