| BIG > Algorithms > ChromosomeJ |
| CONTENTS |
ChromosomeJ is an efficient and user-friendly tool for the segmentation of images containing chromosomes or, more generally, rod-shaped elements, that are "spread" on microscopic slides. The algorithm allows the user to automatically extract the profile of each chromosome and to sort the collection of profiles in a karyotype image (see Figure 1A-B). The method has been incorporated in a plug-in for ImageJ, which is a free open-source multiplatform Java image-processing software. The plug-in is independent of any imaging hardware and, thanks to ImageJ, any common file format may be used as input.
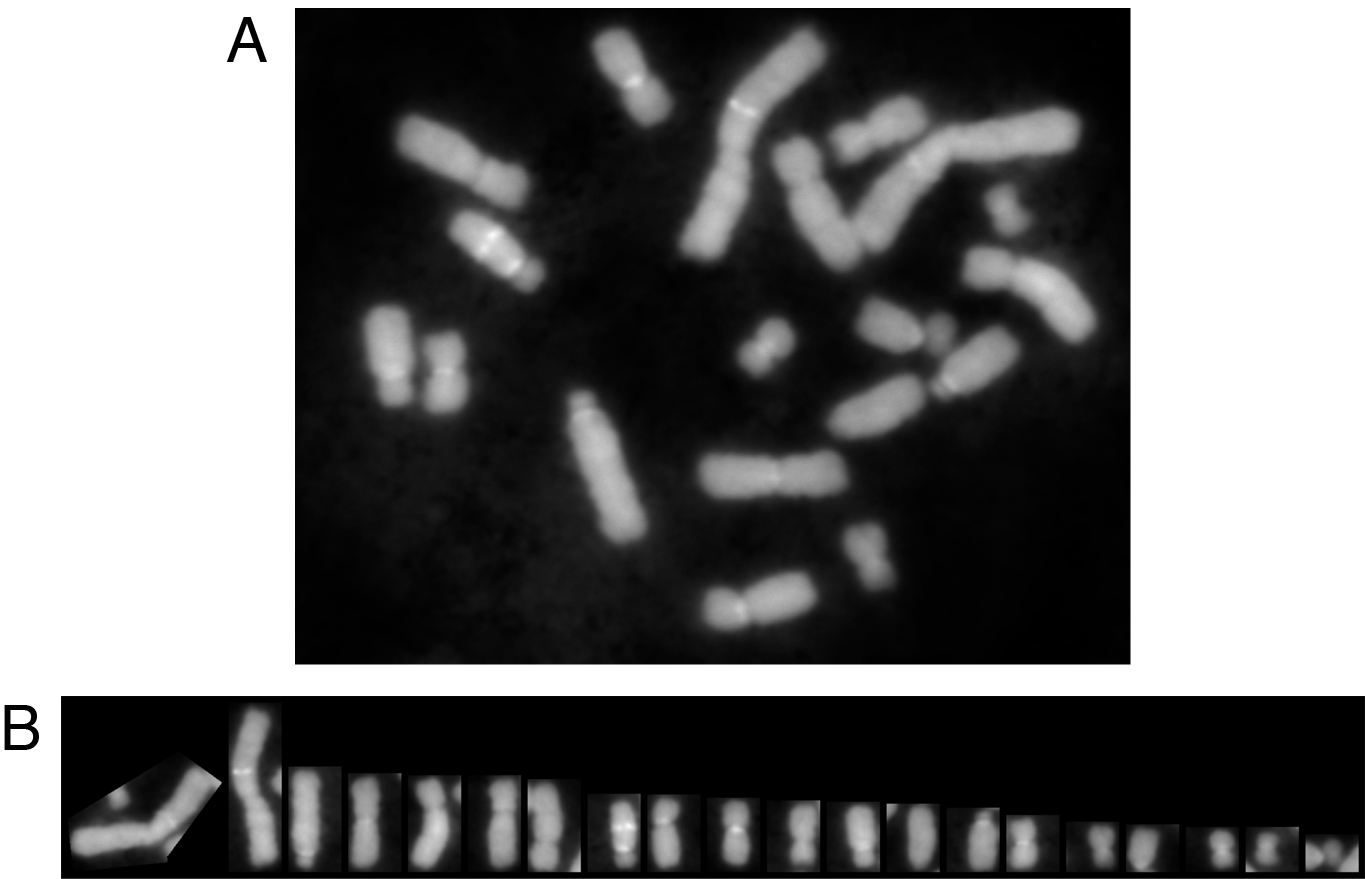
Figure 1. Outline of ChromosomeJ capabilities.
The software provided here is a plugin for ImageJ, a general purpose image-processing and image-analysis package. ImageJ has a public domain licence; it runs on several plateforms: Unix, Linux, Windows, Mac OS 9 and Mac OS X. It doesn't take more than a couple of minutes to install.
To install the plugin download the ChromosomeJ_.jar file and put it in the plugins folder of ImageJ (ImageJ/plugins/). ChromosomeJ also requires some external libraries that have been regrouped in the following archive file. The archive should be uncompressed, and the elements it contains should be placed into the jars folder of the plugins folder of ImageJ (ImageJ/plugins/jars/).
The plugin can be started from the Plugins tab of the ImageJ menu (see Figure 2).
Figure 2. ImageJ plugin menu featuring ChromosomeJ.
The plug-in requires at least one open image at launch. If a rectangular ROI is present in the image, ChromosomeK will automatically crop the image to the ROI and start the analysis on this portion of the data. Otherwise, the entire image will be used.
At launch, the settings window will appear (see Figure 3). This windows allows one to determine several parameters in order to adapt the analysis to the current image. A quick summary of the input parameters is as follows:
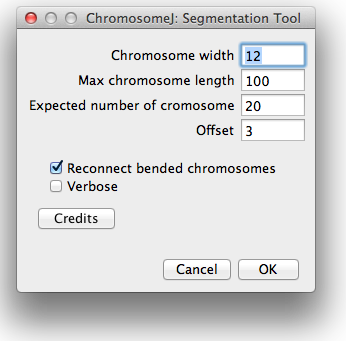
Figure 3. ChromosomeJ settings window.
Using the parameters from the settings window, ChromosomeJ will start an automatic segmentation procedure. If no chromosome is found at the end of this step, a dialog appears and the plugin is closed. If at least one chromosome is found, a validation procedure is launched.
In order to validate automated segmentation results, two windows appear on ImageJ, as shown in Figure 4. On the first one, a summary of all segmentation is shown with multiple rectangles overlaid on the original image. The rectangles are color-coded as follows:
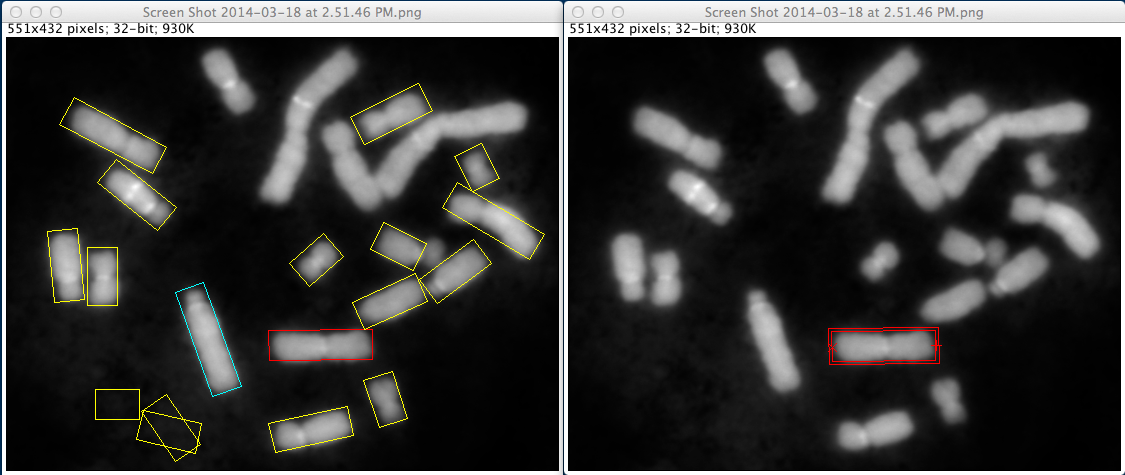
Figure 4. Segmentation validation.
On the second window, the current detection only is overlaid on the original image and the traditional ImageJ toolbar is replaced by a new snake-optimization toolbar (depicted in Figure 5). This allows the user to manually edit one detection at the time.
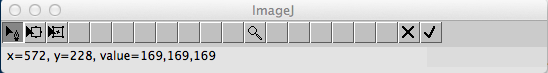
Figure 5. Snake optimization toolbar.
The snake optimization toolbar is composed of several buttons that allow modifying the curve using the control points.
 Move tool: to select a point, click anywhere in the image. The point closest to where you clicked will be highlighted. To move points, drag within the image. The point closest to where you clicked will be moved to the position where you release the mouse. Alternatively, you can use the keyboard arrow keys to get a finer control over the position of the points; for this to work, the image window must be frontmost. Note: it is not possible to interactively move a point outside of the image frame. To translate the snake all at once, you can also depress the SHIFT key before dragging the mouse. Still, a combination of shift and arrow keys will move just one point.
Move tool: to select a point, click anywhere in the image. The point closest to where you clicked will be highlighted. To move points, drag within the image. The point closest to where you clicked will be moved to the position where you release the mouse. Alternatively, you can use the keyboard arrow keys to get a finer control over the position of the points; for this to work, the image window must be frontmost. Note: it is not possible to interactively move a point outside of the image frame. To translate the snake all at once, you can also depress the SHIFT key before dragging the mouse. Still, a combination of shift and arrow keys will move just one point. Scale tool: when this tool is selected, a bounding box appears on the image enclosing the control points of the snake. To move the edges of the bounding box, drag them. The curve will be rescaled accordingly to the new bounding box. To resize the snake all at once, you can also depress the SHIFT key before dragging the mouse.
Scale tool: when this tool is selected, a bounding box appears on the image enclosing the control points of the snake. To move the edges of the bounding box, drag them. The curve will be rescaled accordingly to the new bounding box. To resize the snake all at once, you can also depress the SHIFT key before dragging the mouse. Rotation tool: when this tool is selected, a bounding box appears on the image enclosing the control points of the snake. To rotate the bounding box, drag it. The curve will be rotated accordingly to the rotated bounding box.
Rotation tool: when this tool is selected, a bounding box appears on the image enclosing the control points of the snake. To rotate the bounding box, drag it. The curve will be rotated accordingly to the rotated bounding box. Zoom tool: to magnify the image, select the zoom tool and click in the image. To minify it, use the same tool with the CTRL key or the ALT key.
Zoom tool: to magnify the image, select the zoom tool and click in the image. To minify it, use the same tool with the CTRL key or the ALT key.  Validate tool: this tool is used to accept the detection.
Validate tool: this tool is used to accept the detection. Cancel tool: this tool is used to reject the detection.
Cancel tool: this tool is used to reject the detection.If the appropiate option has been selected from the main menu, when there are no more detections to validate, the second window closes, and a new dialog appears (Figure 6). This dialog allows the user to resolve situations where over-segmentation is observed by informing the plugin that some snakes have captured complementary parts of the same chromosome.
Two rectangle can then be connected as follows:
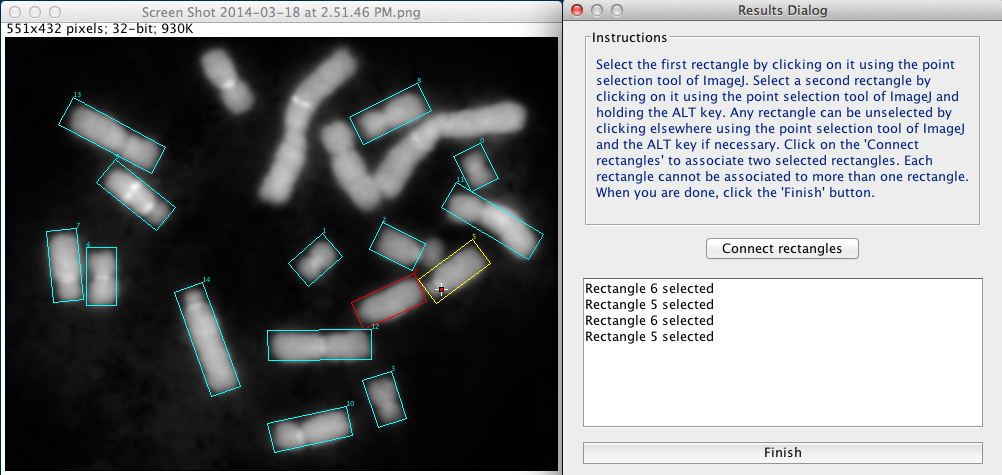
Figure 6. Connecting bended chromosomes.
Once all the required connections have been made between the rectangles, the Finish button can be clicked.
ChromosomeJ finally outputs a new image displaying all the segmented chromosomes sorted by length, as pictured in Figure 7.
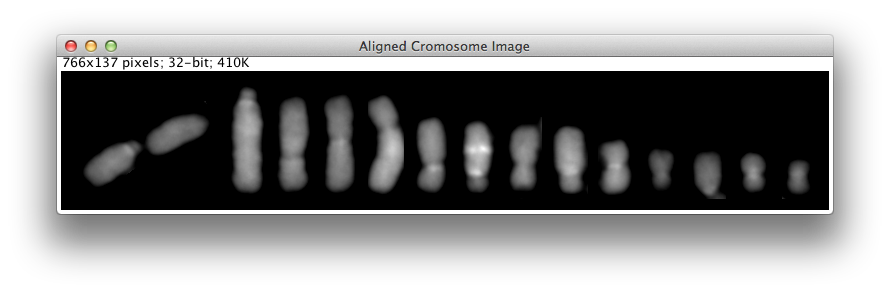
Figure 7. Sample ChromosomeJ output.
You'll be free to use this software for research purposes, but you must not transmit and distribute it without our consent. In addition, you undertake to include a citation to the associated papers whenever you present or publish results that are based on it. EPFL makes no warranties of any kind on this software and shall in no event be liable for damages of any kind in connection with the use and exploitation of this technology.
© 2018 EPFL • webmaster.big@epfl.ch • 09.08.2018