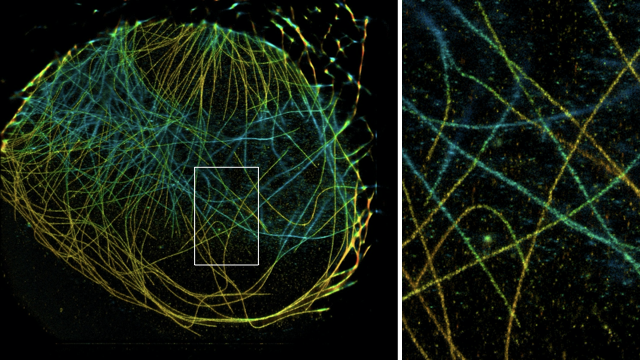
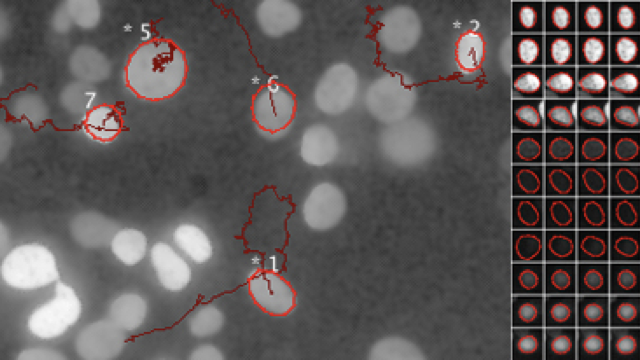
C. Toniolo, D. Sage, J.D. McKinney, N. Dhar,
Quantification of Mycobacterium tuberculosis Growth in Cell-Based Infection Assays by Time-Lapse Fluorescence Microscopy,
Methods in Molecular Biology, vol. 2813, Springer, 2024.
Quantification of Mycobacterium tuberculosis (Mtb) growth dynamics in cell-based in vitro infection models is traditionally carried out by measurement of colony forming units (CFU). However, Mtb being an extremely slow growing organism (16–24 h doubling time), this approach requires at least 3 weeks of incubation to obtain measurable readouts. In this chapter, we describe an alternative approach based on time-lapse microscopy and quantitative image analysis that allows faster quantification of Mtb growth dynamics in host cells. In addition, this approach provides the capability to capture other readouts from the same experimental setup, such as host cell viability, bacterial localization as well as the dynamics of propagation of infection between the host cells.
R. Sarkis, L.-F. Celma, R. Dornier, O. Burri, C. Royer-Chardon, M. Barthélemy, A. Alonso, L. de Leval, D. Sage, O. Naveiras,
MegaQuant: Fully Integrated Deep-Learning Workflow in QuPath—Application to the Detection of Megakaryocytes in Human Bone Marrow,
Proceedings of the European Congress on Digital Pathology (ECDP'24), Vilnius, Lithuania, 2024.
Megakaryocytes (MKs), precursors of platelets, can be altered by clinical conditions and their assessment is of diagnostic value. When evaluated on H&E images, they are challenging to segment due to their complex shape. Here, we propose a fully integrated workflow implemented within QuPath, leveraging existing deep-learning tools to segment and quantify MKs.
C. García-López-de-Haro, S. Dallongeville, T. Musset, E. Gómez-de-Mariscal, D. Sage, W. Ouyang, A. Muñoz-Barrutia, J.-Y. Tinevez, J.-C. Olivo-Marin,
JDLL: A Library to Run Deep Learning Models on Java Bioimage Informatics Platforms,
Nature Methods 21, January 2024.
The advancements in artificial intelligence (AI) technology over the past decade have been a breakthrough in imaging for life sciences, paving the way for novel methods in image restoration, reconstruction and segmentation. However, the wide adoption of deep learning (DL) techniques by end users in bioimage analysis is hindered by the complexity of their deployment. These techniques stem from a variety of rapidly evolving frameworks (for example, TensorFlow 1 or 2, PyTorch) that come with distinct and often conflicting setups, which can discourage even proficient developers. This has led to integration difficulties or even absence in mainstream bioimage informatics platforms such as ImageJ, Icy and Fiji, many of which are primarily developed in Java.
M Eisenmann et al.,
Biomedical image analysis competitions: The state of current participation practice,
arXiv:2212.08568, 2023.
Validation of biomedical image analysis algorithms is typically conducted through so-called challenges - large international benchmarking competitions that compare algorithm performance on identical datasets. Recent years have not only seen an increase in the complexity of the machine learning (ML) models used to solve the tasks, but also a tremendous increase of the scientific impact of challenges, with results often being published in prestigious journals and the winner sometimes receiving important attention in terms of citations and monetary compensa- tion. However, despite this impact, we identified a notable gap in the literature regarding insights into current common practice in challenges. To address this issue, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE In- ternational Symposium on Biomedical Imaging (ISBI) and the International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI) in the year 2021 (80 leaderboards in total). This white paper presents the survey design and the summary of responses.
D.J Nieves, J.A Pike, F. Levet, D. J Williamson, M. Baragilly, S. Oloketuyi, A. de Marco, J. Griffié, D. Sage, E.A.K. Cohen, J.-B. Sibarita, M. Heilemann, D.M. Owen,
A framework for evaluating the performance of SMLM cluster analysis algorithms,
Nature Methods 20, 2023.
Single-molecule localization microscopy (SMLM) generates data in the form of coordinates of localized fluorophores. Cluster analysis is an attractive route for extracting biologically meaningful information from such data and has been widely applied. Despite a range of cluster analysis algorithms, there exists no consensus framework for the evaluation of their performance. Here, we use a systematic approach based on two metrics to score the success of clustering algorithms in simulated conditions mimicking experimental data. We demonstrate the framework using seven diverse analysis algorithms: DBSCAN, ToMATo, KDE, FOCAL, CAML, ClusterViSu and SR-Tesseler. Given that the best performer depended on the underlying distribution of localizations, we demonstrate an analysis pipeline based on statistical similarity measures that enables the selection of the most appropriate algorithm, and the optimized analysis parameters for real SMLM data. We propose that these standard simulated conditions, metrics and analysis pipeline become the basis for future analysis algorithm development and evaluation.
V. Uhlmann, Z. Püspöki, A. Depeursinge, M. Unser, D. Sage, J. Fageot,
Steer’n’Detect: Fast 2D Template Detection with Accurate Orientation Estimation,
Bioinformatics 38, 2022.
Motivation: Rotated template matching is an efficient and versatile algorithm to analyze microscopy images, as it automates the detection of stereotypical structures, such as organelles that can appear at any orientation. Its performance however quickly degrades in noisy image data.
Results: We introduce Steer’n’Detect, an ImageJ plugin implementing a recently published algorithm to detect patterns of interest at any orientation with high accuracy from a single template in 2D images. Steer’n’Detect provides a faster and more robust substitute to template matching. By adapting to the statistics of the image background, it guarantees accurate results even in the presence of noise. The plugin comes with an intuitive user interface facilitating results analysis and further post-processing.
W. Ouyang et al.,
BioImage Model Zoo: A Community-Driven Resource for Accessible Deep Learning in BioImage Analysis,
bioRxiv 2022.06.07.495102, 2022.
Deep learning-based approaches are revolutionizing imaging-driven scientific research. However, the accessibility and reproducibility of deep learning-based workflows for imaging scientists remain far from sufficient. Several tools have recently risen to the challenge of democratizing deep learning by providing user-friendly interfaces to analyze new data with pre-trained or fine-tuned models. Still, few of the existing pre-trained models are interoperable between these tools, critically restricting a model’s overall utility and the possibility of validating and reproducing scientific analyses. Here, we present the BioImage Model Zoo (https://bioimage.io): a community-driven, fully open resource where standardized pre-trained models can be shared, explored, tested, and downloaded for further adaptation or direct deployment in multiple end user-facing tools (e.g., ilastik, deepImageJ, QuPath, StarDist, ImJoy, ZeroCostDL4Mic, CSBDeep). To enable everyone to contribute and consume the Zoo resources, we provide a model standard to enable cross-compatibility, a rich list of example models and practical use-cases, developer tools, documentation, and the accompanying infrastructure for model upload, download and testing. Our contribution aims to lay the groundwork to make deep learning methods for microscopy imaging findable, accessible, interoperable, and reusable (FAIR) across software tools and platforms.
L.G. Jensen, T.Y. Hoh, D.J. Williamson, J. Griffié, D. Sage, P. Rubin-Delanchy, D.M. Owen,
Correction of Multiple-Blinking Artifacts in Photoactivated Localization Microscopy,
Nature Methods 19, 2022.
Photoactivated localization microscopy (PALM) produces an array of localization coordinates by means of photoactivatable fluorescent proteins. However, observations are subject to fluorophore multiple blinking and each protein is included in the dataset an unknown number of times at different positions, due to localization error. This causes artificial clustering to be observed in the data. We present a 'model-based correction' (MBC) workflow using calibration-free estimation of blinking dynamics and model-based clustering to produce a corrected set of localization coordinates representing the true underlying fluorophore locations with enhanced localization precision, outperforming the state of the art. The corrected data can be reliably tested for spatial randomness or analyzed by other clustering approaches, and descriptors such as the absolute number of fluorophores per cluster are now quantifiable, which we validate with simulated data and experimental data with known ground truth. Using MBC, we confirm that the adapter protein, the linker for activation of T cells, is clustered at the T cell immunological synapse.
V. Uhlmann, L. Donati, D. Sage,
A Practical Guide to Supervised Deep Learning for Bioimage Analysis: Challenges and Good Practices,
IEEE Signal Processing Magazine 39, 2022.
The variety of bioimage data and their quality have dramatically increased over the last decade. In parallel, the number of proposed deep learning (DL) models for their analysis grows by the day. Yet, the adequate reuse of published tools by practitioners without DL expertise still raises many practical questions. In this article, we explore four categories of challenges faced by researchers when using supervised DL models in bioimaging applications. We provide examples in which each challenge arises and review the consequences that inadequate decisions may have. We then outline good practices that can be implemented to address the challenges of each category in a scientifically sound way. We provide pointers to the resources that are already available or in active development to help in this endeavor and advocate for the development of further community-driven standards. While primarily intended as a practical tutorial for life scientists, this article also aims at fostering discussions among method developers around the formulation of guidelines for the adequate deployment of DL, with the ultimate goal of accelerating the adoption of novel DL technologies in the biology community.
E. Gómez-de-Mariscal, C. García-López-de-Haro, W. Ouyang, L. Donati, E. Lundberg, M. Unser, A. Muñoz-Barrutia, D. Sage,
DeepImageJ: A User-Friendly Environment to Run Deep Learning Models in ImageJ,
Nature Methods 18, 2021.
DeepImageJ is a user-friendly solution that enables the generic use of pre-trained deep learning models for biomedical image analysis in ImageJ. The deepImageJ environment gives access to the largest bioimage repository of pre-trained deep learning models (BioImage Model Zoo). Hence, nonexperts can easily perform common image processing tasks in life-science research with deep learning-based tools including pixel and object classification, instance segmentation, denoising or virtual staining. DeepImageJ is compatible with existing state of the art solutions and it is equipped with utility tools for developers to include new models. Very recently, several training frameworks have adopted the deepImageJ format to deploy their work in one of the most used softwares in the field (ImageJ). Beyond its direct use, we expect deepImageJ to contribute to the broader dissemination and reuse of deep learning models in life sciences applications and bioimage informatics.
Q. Juppet, F. De Martino, E. Marcandalli, M. Weigert, O. Burri, M. Unser, C. Brisken, D. Sage,
Deep Learning Enables Individual Xenograft Cell Classification in Histological Images by Analysis of Contextual Features,
Journal of Mammary Gland Biology and Neoplasia 26, 2021.
Patient-Derived Xenografts (PDXs) are the preclinical models which best recapitulate inter- and intra-patient complexity of human breast malignancies, and are also emerging as useful tools to study the normal breast epithelium. However, data analysis generated with such models is often confounded by the presence of host cells and can give rise to data misinterpretation. For instance, it is important to discriminate between xenografted and host cells in histological sections prior to performing immunostainings. We developed Single Cell Classifier (SCC), a data-driven deep learning-based computational tool that provides an innovative approach for automated cell species discrimination based on a multi-step process entailing nuclei segmentation and single cell classification. We show that human and murine cell contextual features, more than cell-intrinsic ones, can be exploited to discriminate between cell species in both normal and malignant tissues, yielding up to 96% classification accuracy. SCC will facilitate the interpretation of H&E- and DAPI-stained histological sections of xenografted human-in-mouse tissues and it is open to new in-house built models for further applications. SCC is released as an open-source plugin in ImageJ/Fiji available at the following link: https://github.com/Biomedical-Imaging-Group/SingleCellClassifier.
M. Pourya, S. Aziznejad, M. Unser, D. Sage,
GRAPHIC: Graph-Based Hierarchical Clustering for Single-Molecule Localization Microscopy,
IEEE International Symposium on Biomedical Imaging (ISBI), 2021.
We propose a novel method for the clustering of point-cloud data that originate from single-molecule localization microscopy (SMLM). Our scheme has the ability to infer a hierarchical structure from the data. It takes a particular relevance when quantitatively analyzing the biological particles of interest at different scales. It assumes a prior neither on the shape of particles nor on the background noise. Our multiscale clustering pipeline is built upon graph theory. At each scale, we first construct a weighted graph that represents the SMLM data. Next, we find clusters using spectral clustering. We then use the output of this clustering algorithm to build the graph in the next scale; in this way, we ensure consistency over different scales. We illustrate our method with examples that highlight some of its important properties.
Q. Denoyelle, T.-a. Pham, P. del Aguila Pla, D. Sage, M. Unser,
Optimal-Transport-Based Metric for SMLM,
IEEE International Symposium on Biomedical Imaging (ISBI), 2021.
We propose the use of Flat Metric to assess the performance of reconstruction methods for single-molecule localization microscopy (SMLM) in scenarios where the ground-truth is available. Flat Metric is intimately related to the concept of optimal transport between measures of different mass, providing solid mathematical foundations for SMLM evaluation and integrating both localization and detection performance. In this paper, we provide the foundations of Flat Metric and validate this measure by applying it to controlled synthetic examples and to data from the SMLM 2016 Challenge.
J. Griffié and D. Sage
Virtual-SMLM, a virtual environment for real-time interactive SMLM acquisition,
bioRxiv 2020.03.05.967893, 2020.
Although single molecule localisation microscopy enables for the visualisation of cells nanoscale organisation, its dissemination remains limited mainly due to the complexity of the associated imaging acquisition, impacting on outputs’ reliability and reproducibility. We propose here the first all-in-one fully virtual environment for SMLM acquisition: Virtual-SMLM, including on-the-fly interactivity and real time display. It relies on a novel realistic approach to simulate fluorophores photo-physics based on independent pseudo-continuous emission traces. It also facilitates for user-specific experimental and optical environment design. As such, it constitutes a unique tool for the training of both users and machine learning approaches to automated SMLM, as well as for experimental validation, whilst providing realistic data sets for the development of image reconstruction algorithms and data analysis software.
V. Petrenko, N.R. Gandasi, D. Sage, A. Tengholm, S. Barg, C. Dibner,
In Pancreatic Islets from Type 2 Diabetes Patients, the Dampened Circadian Oscillators Lead to Reduced Insulin and Glucagon Exocytosis,
National Academy of Sciences (PNAS), 117, 2020.
Circadian clocks operative in pancreatic islets participate in the regulation of insulin secretion in humans and, if compromised, in the development of type 2 diabetes (T2D) in rodents. Here we demonstrate that human islet α- and β-cells that bear attenuated clocks exhibit strongly disrupted insulin and glucagon granule docking and exocytosis. To examine whether compromised clocks play a role in the pathogenesis of T2D in humans, we quantified parameters of molecular clocks operative in human T2D islets at population, single islet, and single islet cell levels. Strikingly, our experiments reveal that islets from T2D patients contain clocks with diminished circadian amplitudes and reduced in vitro synchronization capacity compared to their nondiabetic counterparts. Moreover, our data suggest that islet clocks orchestrate temporal profiles of insulin and glucagon secretion in a physiological context. This regulation was disrupted in T2D subjects, implying a role for the islet cell-autonomous clocks in T2D progression. Finally, Nobiletin, an agonist of the core-clock proteins RORα/γ, boosted both circadian amplitude of T2D islet clocks and insulin secretion by these islets. Our study emphasizes a link between the circadian clockwork and T2D and proposes that clock modulators hold promise as putative therapeutic agents for this frequent disorder.
E. Soubies, F. Soulez, M.T. McCann, T.-a. Pham, L. Donati, T. Debarre, D. Sage, M. Unser,
Pocket Guide to Solve Inverse Problems with GlobalBioIm,
Inverse Problems, 35, 2019.
GlobalBioIm is an open-source MATLAB® library for solving inverse problems. The library capitalizes on the strong commonalities between forward models to standardize the resolution of a wide range of imaging inverse problems. Endowed with an operator-algebra mechanism, GlobalBioIm allows one to easily solve inverse problems by combining elementary modules in a lego-like fashion. This user-friendly toolbox gives access to cutting-edge reconstruction algorithms, while its high modularity makes it easily extensible to new modalities and novel reconstruction methods. We expect GlobalBioIm to respond to the needs of imaging scientists looking for reliable and easy-to-use computational tools for solving their inverse problems. In this paper, we present in detail the structure and main features of the library. We also illustrate its flexibility with examples from multichannel deconvolution microscopy.
D. Sage, T.-A. Pham, H. Babcock, T. Lukes, T. Pengo, J. Chao, R. Velmurugan, A. Herbert, A. Agrawal, S. Colabrese, A. Wheeler, A. Archetti, B. Rieger, R. Ober, G.M. Hagen, J.-B. Sibarita, J. Ries, R. Henriques, M. Unser, S. Holden,
Super-Resolution Fight Club: Assessment of 2D and 3D Single-Molecule Localization Microscopy Software,
Nature Methods, 16, 2019.
With the widespread uptake of two-dimensional (2D) and three-dimensional (3D) single-molecule localization microscopy (SMLM), a large set of different data analysis packages have been developed to generate super-resolution images. In a large community effort, we designed a competition to extensively characterize and rank the performance of 2D and 3D SMLM software packages. We generated realistic simulated datasets for popular imaging modalities—2D, astigmatic 3D, biplane 3D and double-helix 3D—and evaluated 36 participant packages against these data. This provides the first broad assessment of 3D SMLM software and provides a holistic view of how the latest 2D and 3D SMLM packages perform in realistic conditions. This resource allows researchers to identify optimal analytical software for their experiments, allows 3D SMLM software developers to benchmark new software against the current state of the art, and provides insight into the current limits of the field.
A. Badoual, A. Galan, D. Sage, M. Unser,
Deforming Tessellations for the Segmentation of Cell Aggregates,
IEEE International Symposium on Biomedical Imaging (ISBI), 2019.
We present a new active contour to segment cell aggregates. We describe it by a smooth tessellation that is attracted toward the cell membranes. Our approach relies on subdivision schemes that are tightly linked to the theory of wavelets. The shape is encoded by control points grouped in tiles. The smooth and continuously defined boundary of each tile is generated by recursively applying a refinement process to its control points. We deform the smooth tessellation in a global manner using a ridge-based energy that we have designed for that purpose. By construction, cells are segmented without overlap and the tessellation structure is maintained even on dim membranes. Leakage, which afflicts usual image-processing methods (e.g., watershed), is thus prevented. We validate our framework on both synthetic and real microscopy images, showing that the proposed method is robust to membrane gaps and to high levels of noise.
T.-a. Pham, E. Soubies, D. Sage, M. Unser,
Closed-Form Expression of the Fourier Ring-Correlation for Single-Molecule Localization Microscopy,
Best student paper award, IEEE International Symposium on Biomedical Imaging (ISBI), 2019.
Single-molecule localization microscopy (SMLM) is a popular microscopic technique that achieves super resolution imaging by localizing individual blinking molecules in thousands of frames. Therefore, the reconstructed high-resolution image is a combination of millions of point sources. This particular computational reconstruction leads to the question of the estimation of the image resolution. Fourier-ring correlation (FRC) is the standard tool for assessing the resolution. It has been proposed for SMLM by computing a discrete correlation in the Fourier domain. In this work, we derive a closed-form expression to compute the continuous FRC. Our implementation provides an exact FRC and an alternative to compute a parameter-free FRC. In addition, it gives insights on the discrepancy of the discrete FRC and yields a rule to select its parameters such as the spatial sampling step or the width of the kernel used as density estimator.
A.I. Rahaghi, U. Lemmin, D. Sage, D.A. Barry,
Achieving High-Resolution Thermal Imagery in Low-Contrast Lake Surface Waters by Aerial Remote Sensing and Image Registration,
Remote Sensing of Environment, 221, 2019.
A two-platform measurement system for realizing airborne thermography of the Lake Surface Water Temperature (LSWT) with ~0.8m pixel resolution (sub-pixel satellite scale) is presented. It consists of a tethered Balloon Launched Imaging and Monitoring Platform (BLIMP) that records LSWT images and an autonomously operating catamaran (called ZiviCat) that measures in situ surface/near surface temperatures within the image area, thus permitting simultaneous ground-truthing of the BLIMP data. The BLIMP was equipped with an uncooled InfraRed (IR) camera. The ZiviCat was designed to measure along predefined trajectories on a lake. Since LSWT spatial variability in each image is expected to be low, a poor estimation of the common spatial and temporal noise of the IR camera (nonuniformity and shutter-based drift, respectively) leads to errors in the thermal maps obtained. Nonuniformity was corrected by applying a pixelwise two-point linear correction method based on laboratory experiments. A Probability Density Function (PDF) matching in regions of overlap between sequential images was used for the drift correction. A feature matching-based algorithm, combining blob and region detectors, was implemented to create composite thermal images, and a mean value of the overlapped images at each location was considered as a representative value of that pixel in the final map. The results indicate that a high overlapping field of view (~95%) is essential for image fusion and noise reduction over such low-contrast scenes. The in situ temperatures measured by the ZiviCat were then used for the radiometric calibration. This resulted in the generation of LSWT maps at sub-pixel satellite scale resolution that revealed spatial LSWT variability, organized in narrow streaks hundreds of meters long and coherent patches of different size, with unprecedented detail.
H. Deschout, I. Platzman, D. Sage, L. Feletti, J.P. Spatz, A. Radenovic,
Investigating Focal Adhesion Substructures by Localization Microscopy,
Biophysical Journal 113, 2017.
Cells rely on focal adhesions (FAs) to carry out a variety of important tasks, including motion, environmental sensing, and adhesion to the extracellular matrix. Although attaining a fundamental characterization of FAs is a compelling goal, their extensive complexity and small size, which can be below the diffraction limit, have hindered a full understanding. In this study we have used single-molecule localization microscopy (SMLM) to investigate integrin β3 and paxillin in rat embryonic fibroblasts growing on two different extracellular matrix-representing substrates (i.e., fibronectin-coated substrates and specifically biofunctionalized nanopatterned substrates). To quantify the substructure of FAs, we developed a clustering method based on expectation maximization of a Gaussian mixture that accounts for localization uncertainty and background. Analysis of our SMLM data indicates that the structures within FAs, characterized as a Gaussian mixture, typically have areas between 0.01 and 1 μm2, contain 10–100 localizations, and can exhibit substantial eccentricity. Our approach based on SMLM opens new avenues for studying structural and functional biology of molecular assemblies that display substantial varieties in size, shape, and density.
R. Schorer, A. Badoual, B. Bastide, A. Vandebrouck, M. Licker, D. Sage,
A Feasability Study of Color Flow Doppler Vectorization for Automated Blood Flow Monitoring,
Journal of Clinical Monitoring and Computing 31, 2017.
An ongoing issue in vascular medicine is the measure of the blood flow. Catheterization remains the gold standard measurement method, although non-invasive techniques are an area of intense research. We hereby present a computational method for real-time measurement of the blood flow from color flow Doppler data, with a focus on simplicity and monitoring instead of diagnostics. We then analyze the performance of a proof-of-principle software implementation. We imagined a geometrical model geared towards blood flow computation from a color flow Doppler signal, and we developed a software implementation requiring only a standard diagnostic ultrasound device. Detection performance was evaluated by computing flow and its determinants (flow speed, vessel area, and ultrasound beam angle of incidence) on purposely designed synthetic and phantom-based arterial flow simulations. Flow was appropriately detected in all cases. Errors on synthetic images ranged from nonexistent to substantial depending on experimental conditions. Mean errors on measurements from our phantom flow simulation ranged from 1.2 to 40.2% for angle estimation, and from 3.2 to 25.3% for real-time flow estimation. This study is a proof of concept showing that accurate measurement can be done from automated color flow Doppler signal extraction, providing the industry the opportunity for further optimization using raw ultrasound data.
V. Petrenko, C. Saini, L. Giovannoni, C. Gobet, D. Sage, M. Unser, M.H. Masson, G. Gu, D. Bosco, F. Gachon, J. Philippe, C. Dibner,
Pancreatic α- and β-Cellular Clocks Have Distinct Molecular Properties and Impact on Islet Hormone Secretion and Gene Expression,
Genes & Development 31, 2017.
A critical role of circadian oscillators in orchestrating insulin secretion and islet gene transcription has been demonstrated recently. However, these studies focused on whole islets and did not explore the interplay between α-cell and β-cell clocks. We performed a parallel analysis of the molecular properties of α-cell and β-cell oscillators using a mouse model expressing three reporter genes: one labeling α cells, one specific for β cells, and a third monitoring circadian gene expression. Thus, phase entrainment properties, gene expression, and functional outputs of the α-cell and β-cell clockworks could be assessed in vivo and in vitro at the population and single-cell level. These experiments showed that α-cellular and β-cellular clocks are oscillating with distinct phases in vivo and in vitro. Diurnal transcriptome analysis in separated α and β cells revealed that a high number of genes with key roles in islet physiology, including regulators of glucose sensing and hormone secretion, are differentially expressed in these cell types. Moreover, temporal insulin and glucagon secretion exhibited distinct oscillatory profiles both in vivo and in vitro. Altogether, our data indicate that differential entrainment characteristics of circadian α-cell and β-cell clocks are an important feature in the temporal coordination of endocrine function and gene expression.
D. Sage, L. Donati, F. Soulez, D. Fortun, G. Schmit, A. Seitz, R. Guiet, C. Vonesch, M. Unser,
DeconvolutionLab2: An Open-Source Software for Deconvolution Microscopy,
Methods—Image Processing for Biologists 115, 2017.
Images in fluorescence microscopy are inherently blurred due to the limit of diffraction of light. The purpose of deconvolution microscopy is to compensate numerically for this degradation. Deconvolution is widely used to restore fine details of 3D biological samples. Unfortunately, dealing with deconvolution tools is not straightforward. Among others, end users have to select the appropriate algorithm, calibration and parametrization, while potentially facing demanding computational tasks. To make deconvolution more accessible, we have developed a practical platform for deconvolution microscopy called DeconvolutionLab. Freely distributed, DeconvolutionLab hosts standard algorithms for 3D microscopy deconvolution and drives them through a user-oriented interface. In this paper, we take advantage of the release of DeconvolutionLab2 to provide a complete description of the software package and its built-in deconvolution algorithms. We examine several standard algorithms used in deconvolution microscopy, notably: Regularized inverse filter, Tikhonov regularization, Landweber, Tikhonov-Miller, Richardson-Lucy, and fast iterative shrinkage-thresholding. We evaluate these methods over large 3D microscopy images using simulated datasets and real experimental images. We distinguish the algorithms in terms of image quality, performance, usability and computational requirements. Our presentation is completed with a discussion of recent trends in deconvolution, inspired by the results of the Grand Challenge on deconvolution microscopy that was recently organized.
K. Wasmer, P.-M. Pochon, D. Sage, J.H. Giovanola,
Parametric Experimental Study and Design of Experiment Modelling of Sapphire Grinding,
Journal of Cleaner Production 141, 2017.
This study investigates and models the grinding process of single crystal sapphire. Five parameters: the wheel speed, the feed speed, the vertical feed, the ultrasonic assistance and the crystallographic direction were considered via a design of experiments (DoE) approach. The responses were multiple but can be divided in three groups: the process, the machine and the grinding quality. DoE results revealed that the parameters interact in a complex manner and depends on the responses. Therefore, to gain a better understanding of the grinding process of sapphire, the interactions between parameters have also to be taken into consideration. It was found that three main parameters have the largest influences on the tangential grinding forces: the wheel speed, the feed speed and the vertical feed. In contrast, the median defect area is mainly impacted by the quadratic effects of the wheel speed and vertical feed followed by various interactions. After an optimization procedure, the second optimum for the tangential forces was found to be very close to the best optimum for the median defect area. The optimum solution is: a wheel speed of 7'500 rpm, a feed speed of 60 mm∕min, a vertical feed of 12.5 μm∕pass, no ultrasonic assistance and grinding along the c-axis. This set of parameters was validated with additional and repeated tests on both Verneuil and Kyropouloas sapphire. Finally, it came out that the optimum solution has also a very good productivity.
B. Meylan, P. Dogan, D. Sage, K. Wasmer,
A Simple, Fast and Low-Cost Method for in situ Monitoring of Topographical Changes and Wear Rate of a Complex Tribo-System Under Mixed Lubrication,
Wear—An International Journal on the Science and Technology of Friction, 2016.
This contribution presents a simple, fast, and low-cost method to track in situ and in real time the evolution of the surface topography. The method combines an optical method (collimated light) with image analysis. The method was validated using a complex tribo-system; a reciprocal sliding of a rough cast iron counter-piece under mixed lubrication. We demonstrated that the optical method is well suited to observe the evolution of the contact areas during sliding. We also proved that the contact area occurs on the highest peaks of the surface roughness. Finally, we estimated the wear rate by combining the information of the contact area and the Abbott-Firestone curve obtained by profilometry. The wear rate was found to be 40 times higher in the early stage of sliding as compared to the steady-state. The running-in of this particular system was found to be approximately 10 h. In addition to be simple, fast and low-cost, the proposed method has other advantages. It is adaptable for real industrial tests conditions. The fast data analysis allows the wear rate of a tribo-system being determined in real-time. This method characterise the real contact areas of the observed piece, which is difficult, if not impossible, to obtain by profilometric observation alone.
Z. Püspöki, D. Sage, J.P. Ward, M. Unser,
SpotCaliper: Fast Wavelet-Based Spot Detection with Accurate Size Estimation,
Bioinformatics 32, 2016.
Motivation: SpotCaliper is a novel wavelet-based image-analysis software providing a fast automatic detection scheme for circular patterns (spots), combined with the precise estimation of their size. It is implemented as an ImageJ plugin with a friendly user interface. The user is allowed to edit the results by modifying the measurements (in a semi-automated way), extract data for further analysis. The fine tuning of the detections includes the possibility of adjusting or removing the original detections, as well as adding further spots. Results: The main advantage of the software is its ability to capture the size of spots in a fast and accurate way. Availability and implementation: http://bigwww.epfl.ch/algorithms/spotcaliper/ Contact: zsuzsanna.puspoki@epfl.ch Supplementary information: Supplementary data are available at Bioinformatics online.
S. Holden, D. Sage,
Super-Resolution Fight Club,
Nature Photonics 10, 2016.
A 2016 competition to find the optimum software for 3D single-molecule localization microscopy will help practitioners choose the best tool for the job and spur further developments in the field.
E. Bostan, E. Froustey, M. Nilchian, D. Sage, M. Unser,
Variational Phase Imaging Using the Transport-of-Intensity Equation,
IEEE Transactions on Image Processing 25, 2016.
We introduce a variational phase retrieval algorithm for the imaging of transparent objects. Our formalism is based on the transport-of-intensity equation (TIE), which relates the phase of an optical field to the variation of its intensity along the direction of propagation. TIE practically requires one to record a set of defocus images to measure the variation of intensity. We first investigate the effect of the defocus distance on the retrieved phase map. Based on our analysis, we propose a weighted phase reconstruction algorithm yielding a phase map that minimizes a convex functional. The method is nonlinear and combines different ranges of spatial frequencies—depending on the defocus value of the measurements—in a regularized fashion. The minimization task is solved iteratively via the alternating-direction method of multipliers. Our simulations outperform commonly used linear and nonlinear TIE solvers. We also illustrate and validate our method on real microscopy data of HeLa cells.
Z. Püspöki, J.P. Ward, D. Sage, M. Unser,
On the Continuous Steering of the Scale of Tight Wavelet Frames,
SIAM Journal on Imaging Sciences 9, 2016.
In analogy with steerable wavelets, we present a general construction of adaptable tight wavelet frames, with an emphasis on scaling operations. In particular, the derived wavelets can be "dilated" by a procedure comparable to the operation of steering steerable wavelets. The fundamental aspects of the construction are the same: an admissible collection of Fourier multipliers is used to extend a tight wavelet frame, and the "scale" of the wavelets is adapted by scaling the multipliers. As an application, the proposed wavelets can be used to improve the frequency localization. Importantly, the localized frequency bands specified by this construction can be scaled efficiently using matrix multiplication.
Z. Püspöki, M. Storath, D. Sage, M. Unser,
Transforms and Operators for Directional Bioimage Analysis: A Survey,
Focus on Bio-Image Informatics, Springer, 2016.
We give a methodology-oriented perspective on directional image analysis and rotation-invariant processing. We review the state of the art in the field and make connections with recent mathematical developments in functional analysis and wavelet theory. We unify our perspective within a common framework using operators. The intent is to provide image-processing methods that can be deployed in algorithms that analyze biomedical images with improved rotation invariance and high directional sensitivity. We start our survey with classical methods such as directional-gradient and the structure tensor. Then, we discuss how these methods can be improved with respect to robustness, invariance to geometric transformations (with a particular interest in scaling), and computation cost. To address robustness against noise, we move forward to higher degrees of directional selectivity and discuss Hessian-based detection schemes. To present multiscale approaches, we explain the differences between Fourier filters, directional wavelets, curvelets, and shearlets. To reduce the computational cost, we address the problem of matching directional patterns by proposing steerable filters, where one might perform arbitrary rotations and optimizations without discretizing the orientation. We define the property of steerability and give an introduction to the design of steerable filters. We cover the spectrum from simple steerable filters through pyramid schemes up to steerable wavelets. We also present illustrations on the design of steerable wavelets and their application to pattern recognition.
D. Sage, H. Kirshner, T. Pengo, N. Stuurman, J. Min, S. Manley, M. Unser,
Quantitative Evaluation of Software Packages for Single-Molecule Localization Microscopy,
Nature Methods 12, 2015.
The quality of super-resolution images obtained by single-molecule localization microscopy (SMLM) depends largely on the software used to detect and accurately localize point sources. In this work, we focus on the computational aspects of super-resolution microscopy and present a comprehensive evaluation of localization software packages. Our philosophy is to evaluate each package as a whole, thus maintaining the integrity of the software. We prepared synthetic data that represent three-dimensional structures modeled after biological components, taking excitation parameters, noise sources, point-spread functions and pixelation into account. We then asked developers to run their software on our data; most responded favorably, allowing us to present a broad picture of the methods available. We evaluated their results using quantitative and user-interpretable criteria: detection rate, accuracy, quality of image reconstruction, resolution, software usability and computational resources. These metrics reflect the various tradeoffs of SMLM software packages and help users to choose the software that fits their needs.
Z. Püspöki, J.P. Ward, D. Sage, M. Unser,
Fast Detection and Refined Scale Estimation Using Complex Isotropic Wavelets,
IEEE International Symposium on Biomedical Imaging (ISBI), 2015.
The dyadic scaling in the discrete wavelet transform can lead to a loss of precision, in comparison to the computationally unrealistic continuous wavelet transform. To overcome this obstacle, we propose a novel method to locally scale wavelets between dyadic scales in an efficient way. We compute complex wavelet coefficients for a tight frame with a dyadic scale progression. Our isotropic complex wavelets are designed such that the deviation from the nominal scale is encoded in the phase of the coefficients. Moreover, the magnitude of the coefficients is used for feature detection. Numerical experiments are presented to justify our method, and we present results for feature extraction from real data.
P. Wachowicz, A. Chasapi, A. Krapp, E.C. del Rosario, D. Schmitter, D. Sage, M. Unser, I. Xenarios, J. Rougemont, V. Simanis,
Analysis of S. pombe SIN Protein Association to the SPB Reveals Two Genetically Separable States of the SIN,
Journal of Cell Science 128, 2015.
The Schizosaccharomyces pombe septation initiation network (SIN) regulates cytokinesis, and asymmetric association of SIN proteins with the mitotic spindle pole bodies (SPBs) is important for its regulation. Here, we have used semi-automated image analysis to study SIN proteins in large numbers of wild-type and mutant cells. Our principal conclusions are: first, that the association of Cdc7p with the SPBs in early mitosis is frequently asymmetric, with a bias in favour of the new SPB; second, that the early association of Cdc7p-GFP to the SPB depends on Plo1p but not Spg1p, and is unaffected by mutations that influence its asymmetry in anaphase; third, that Cdc7p asymmetry in anaphase B is delayed by Pom1p and by activation of the spindle assembly checkpoint, and is promoted by Rad24p; and fourth, that the length of the spindle, expressed as a fraction of the length of the cell, at which Cdc7p becomes asymmetric is similar in cells dividing at different sizes. These data reveal that multiple regulatory mechanisms control the SIN in mitosis and lead us to propose a two-state model to describe the SIN.
E. Bostan, E. Froustey, B. Rappaz, É. Shaffer, D. Sage, M. Unser,
Phase Retrieval by Using Transport-of-Intensity Equation and Differential Interference Contrast Microscopy,
2014 IEEE International Conference on Image Processing (ICIP), 2014.
We present a variational reconstruction algorithm for the phase-retrieval problem by using the differential interference contrast microscopy. Principally, we rely on the transport-of-intensity equation that specifies the sought phase as the solution of a partial differential equation. Our approach is based on an iterative reconstruction algorithm involving the total variation regularisation which is efficiently solved via the alternating direction method of multipliers. We illustrate the applicability of the method via real data experiments. To the best of our knowledge, this work demonstrates the performance of such an iterative algorithm on real data for the first time.
D. Wüstner, T. Christensen, L.M. Solanko, D. Sage,
Photobleaching Kinetics and Time-Integrated Emission of Fluorescent Probes in Cellular Membranes,
Molecules 19, 2014.
Since the pioneering work of Hirschfeld, it is known that time-integrated emission (TiEm) of a fluorophore is independent of fluorescence quantum yield and illumination intensity. Practical implementation of this important result for determining exact probe distribution in living cells is often hampered by the presence of autofluorescence. Using kinetic modelling of photobleaching combined with pixel-wise bleach rate fitting of decay models with an updated plugin to the ImageJ program, it is shown that the TiEm of a fluorophore in living cells can be determined exactly from the product of bleaching amplitude and time constant. This applies to mono-exponential bleaching from the first excited singlet and/or triplet state and to multi-exponential combinations of such processes. The TiEm can be used to correct for illumination shading and background autofluorescence without the need for fluorescent test layers or separate imaging of non-stained cells. We apply the method to simulated images and to images of cells, whose membranes were labelled with fluorescent sterols and sphingolipids. Our bleaching model can be extended to include a probability density function (PDF) of intrinsic bleach rate constants with a memory kernel. This approach results in a time-dependent bleach rate coefficient and is exemplified for fluorescent sterols in restricted intracellular environments, like lipid droplets. We show that for small deviations from the classical exponential bleaching, the TiEm of decay functions with rate coefficients remains largely independent of fluorescence lifetime and illumination, and thereby represents a faithful measure of probe distribution.
T. Mannic, P. Meyer, F. Triponez, M. Pusztaszeri, G. Le Martelot, O. Mariani, D. Schmitter, D. Sage, J. Philippe, C. Dibner,
Circadian Clock Characteristics Are Altered in Human Thyroid Malignant Nodules,
The Journal of Clinical Endocrinology & Metabolism 98, 2013.
Context: The circadian clock represents the body's molecular time-keeping system. Recent findings revealed strong changes of clock gene expression in various types of human cancers. Objective: Due to emerging evidence on the connection between the circadian oscillator, cell cycle, and oncogenic transformation, we aimed to characterize the circadian clockwork in human benign and malignant thyroid nodules. Design: Clock transcript levels were assessed by quantitative RT-PCR in thyroid tissues. To provide molecular characteristics of human thyroid clockwork, primary thyrocytes established from normal or nodular thyroid tissue biopsies were subjected to in vitro synchronization with subsequent clock gene expression analysis by circadian bioluminescence reporter assay and by quantitative RT-PCR. Results: The expression levels of the Bmal1 were up-regulated in tissue samples of follicular thyroid carcinoma (FTC), and in papillary thyroid carcinoma (PTC), as compared with normal thyroid and benign nodules, whereas Cry2 was down-regulated in FTCandPTC. Human thyrocytes derived from normal thyroid tissue exhibited high-amplitude circadian oscillations of Bmal1-luciferase reporter expression and endogenous clock transcripts. Thyrocytes established from FTC and PTC exhibited clock transcript oscillations similar to those of normal thyroid tissue and benign nodules (except for Per2 altered in PTC), whereas cells derived from poorly differentiated thyroid carcinoma exhibited altered circadian oscillations. Conclusions: This is the first study demonstrating a molecular makeup of the human thyroid circadian clock. Characterization of the thyroid clock machinery alterations upon thyroid nodule malignant transformation contributes to understanding the connections between circadian clocks and oncogenic transformation. Moreover, it might help in improving the thyroid nodule preoperative diagnostics.
D. Sage, H. Kirshner, C. Vonesch, S. Lefkimmiatis, M. Unser,
Benchmarking Image-Processing Algorithms for Biomicroscopy: Reference Datasets and Perspectives,
European Signal Processing Conference (EUSIPCO'13), Marrakech, Morocco, September 9-13, 2013.
As the field of bioimage informatics matures, the issue of the validation of image reconstruction algorithms and the definition of proper performance criteria becomes more pressing. In this work, we discuss benchmarking aspects of fluorescence microscopy quantitative tools. We point out the importance of generating realistic datasets and describe our approach to this task. We rely on our experience and present arguments in favor of the use of 3D continuous-domain models of biological structures for simulating bioimaging datasets. We also present physically-realistic models of image formation that that are reasonably efficiently to implement.
M. Unser, D. Sage, R. Delgado-Gonzalo,
Advanced Image Processing for Biology, and the Open Bio Image Alliance (OBIA),
European Signal Processing Conference (EUSIPCO), Marrakech, Morocco, 2013.
The field of biological imaging has evolved considerably during the past decade as a result of recent (r)evolutions in fluorescence labeling and optical microscopy. Bioimage informatics has been identified as a top priority to cope with the ever-increasing amount of microscopy data. The challenges and opportunities for researchers in image and signal processing are manyfold. They span the areas of mathematical imaging, with problems such as denoising, 3-D deconvolution and super-resolution localization, as well as image analysis for the segmentation, detection and recognition of biological structures in 3-D. The dynamic aspect of the data requires the development of novel algorithms for tracking fluorescent particles and analyzing high-throughput microscopy data (labeling of cells, phenotyping, extraction of gene expression profiles). A crucial aspect of bioimage informatics is making image analysis tools available to biologists so that they can be applied to real data and used on a routine basis. Developers may benefit from open-source frameworks and international initiative such as OBIA for easying-up this process and creating collaboration networks with biologists.
D. Schmitter, P. Wachowicz, D. Sage, A. Chasapi, I. Xenarios, V. Simanis, M. Unser,
A 2D/3D Image Analysis System to Track Fluorescently Labeled Structures in Rod-Shaped Cells: Application to Measure Spindle Pole Asymmetry During Mitosis,
Cell Division 8, 2013.
Background: The yeast Schizosaccharomyces pombe is frequently used as a model for studying the cell cycle. The cells are rod-shaped and divide by medial fission. The process of cell division, or cytokinesis, is controlled by a network of signaling proteins called the Septation Initiation Network (SIN); SIN proteins associate with the SPBs during nuclear division (mitosis). Some SIN proteins associate with both SPBs early in mitosis, and then display strongly asymmetric signal intensity at the SPBs in late mitosis, just before cytokinesis. This asymmetry is thought to be important for correct regulation of SIN signaling, and coordination of cytokinesis and mitosis. In order to study the dynamics of organelles or large protein complexes such as the spindle pole body (SPB), which have been labeled with a fluorescent protein tag in living cells, a number of the image analysis problems must be solved; the cell outline must be detected automatically, and the position and signal intensity associated with the structures of interest within the cell must be determined. Results: We present a new 2D and 3D image analysis system that permits versatile and robust analysis of motile, fluorescently labeled structures in rod-shaped cells. We have designed an image analysis system that we have implemented as a user-friendly software package allowing the fast and robust image-analysis of large numbers of rod-shaped cells. We have developed new robust algorithms, which we combined with existing methodologies to facilitate fast and accurate analysis. Our software permits the detection and segmentation of rod-shaped cells in either static or dynamic (i.e. time lapse) multi-channel images. It enables tracking of two structures (for example SPBs) in two different image channels. For 2D or 3D static images, the locations of the structures are identified, and then intensity values are extracted together with several quantitative parameters, such as length, width, cell orientation, background fluorescence and the distance between the structures of interest. Furthermore, two kinds of kymographs of the tracked structures can be established, one representing the migration with respect to their relative position, the other representing their individual trajectories inside the cell. This software package, called "RodCellJ", allowed us to analyze a large number of S. pombe cells to understand the rules that govern SIN protein asymmetry. Conclusions: "RodCellJ" is freely available to the community as a package of several ImageJ plugins to simultaneously analyze the behavior of a large number of rod-shaped cells in an extensive manner. The integration of different image-processing techniques in a single package, as well as the development of novel algorithms does not only allow to speed up the analysis with respect to the usage of existing tools, but also accounts for higher accuracy. Its utility was demonstrated on both 2D and 3D static and dynamic images to study the septation initiation network of the yeast Schizosaccharomyces pombe. More generally, it can be used in any kind of biological context where fluorescent-protein labeled structures need to be analyzed in rod-shaped cells. Availability: RodCellJ is freely available under http://bigwww.epfl.ch/algorithms.html.
P. Pulimeno, T. Mannic, D. Sage, L. Giovannoni, P. Salmon, S. Lemeille, M. Giry-Laterrière, M. Unser, D. Bosco, C. Bauer, J. Morf, P. Halban, J. Philippe, C. Dibner,
Autonomous and Self-Sustained Circadian Oscillators Displayed in Human Islet Cells,
Diabetologia, 56, 2013.
Aim/hypothesis: Following on from the emerging importance of the pancreas circadian clock on islet function and the development of type 2 diabetes in rodent models, we aimed to examine circadian gene expression in human islets. The oscillator properties were assessed in intact islets as well as in beta cells. Methods: We established a system for long-term bioluminescence recording in cultured human islets, employing lentivector gene delivery of the core clock gene Bmal1 (also known as Arntl)-luciferase reporter. Beta cells were stably labelled using a rat insulin2 promoter fluorescent construct. Single-islet/cell oscillation profiles were measured by combined bioluminescence-fluorescence time-lapse microscopy. Results: Human islets synchronised in vitro exhibited self-sustained circadian oscillations of Bmal1-luciferase expression at both the population and single-islet levels, with period lengths of 23.6 and 23.9 h, respectively. Endogenous BMAL1 and CRY1 transcript expression was circadian in synchronised islets over 48 h, and antiphasic to REV-ERBα (also known as NR1D1), PER1, PER2, PER3 and DBP transcript circadian profiles. HNF1A and PDX1 exhibited weak circadian oscillations, in phase with the REV-ERBα transcript. Dispersed islet cells were strongly oscillating as well, at population and single-cell levels. Importantly, beta and non-beta cells revealed oscillatory profiles that were well synchronised with each other. Conclusions/interpretation: We provide for the first time compelling evidence for high-amplitude cell-autonomous circadian oscillators displayed in human pancreatic islets and in dispersed human islet cells. Moreover, these clocks are synchronised between beta and non-beta cells in primary human islet cell cultures.
M. Roccio, D. Schmitter, M. Knobloch, Y. Okawa, D. Sage, M.P. Lutolf,
Predicting Stem Cell Fate Changes by Differential Cell Cycle Progression Patterns,
Development, 140, 2013.
Stem cell self-renewal, commitment and reprogramming rely on a poorly understood coordination of cell cycle progression and execution of cell fate choices. Using existing experimental paradigms, it has not been possible to probe this relationship systematically in live stem cells in vitro or in vivo. Alterations in stem cell cycle kinetics probably occur long before changes in phenotypic markers are apparent and could be used as predictive parameters to reveal changes in stem cell fate. To explore this intriguing concept, we developed a single-cell tracking approach that enables automatic detection of cell cycle phases in live (stem) cells expressing fluorescent ubiquitylation-based cell-cycle indicator (FUCCI) probes. Using this tool, we have identified distinctive changes in lengths and fluorescence intensities of G1 (red fluorescence) and S/G2-M (green) that are associated with self-renewal and differentiation of single murine neural stem/progenitor cells (NSCs) and embryonic stem cells (ESCs). We further exploited these distinctive features using fluorescence-activated cell sorting to select for desired stem cell fates in two challenging cell culture settings. First, as G1 length was found to nearly double during NSC differentiation, resulting in progressively increasing red fluorescence intensity, we successfully purified stem cells from heterogeneous cell populations by their lower fluorescence. Second, as ESCs are almost exclusively marked by the green (S/G2-M) FUCCI probe due to their very short G1, we substantially augmented the proportion of reprogramming cells by sorting green cells early on during reprogramming from a NSC to an induced pluripotent stem cell state. Taken together, our studies begin to shed light on the crucial relationship between cell cycle progression and fate choice, and we are convinced that the presented approach can be exploited to predict and manipulate cell fate in a wealth of other mammalian cell systems.
H. Kirshner, F. Aguet, D. Sage, M. Unser,
3-D PSF Fitting for Fluorescence Microscopy: Implementation and Localization Application,
Journal of Microscopy 249, 2013.
Localization microscopy relies on computationally efficient Gaussian approximations of the point spread function for the calculation of fluorophore positions. Theoretical predictions show that under specific experimental conditions, localization accuracy is significantly improved when the localization is performed using a more realistic model. Here, we show how this can be achieved by considering three-dimensional (3-D) point spread function models for the wide field microscope. We introduce a least-squares point spread function fitting framework that utilizes the Gibson and Lanni model and propose a computationally efficient way for evaluating its derivative functions. We demonstrate the usefulness of the proposed approach with algorithms for particle localization and defocus estimation, both implemented as plugins for ImageJ.
D. Wüstner, L.M. Solanko, F.W. Lund, D. Sage, H.J. Schroll, M.A. Lomholt,
Quantitative Fluorescence Loss in Photobleaching for Analysis of Protein Transport and Aggregation,
BMC Bioinformatics 13, 2012.
Background
P. Thévenaz, D. Sage, M. Unser,
Bi-Exponential Edge-Preserving Smoother,
IEEE Transactions on Image Processing, 21, 2012.
Edge-preserving smoothers need not be taxed by a severe computational cost. We present in this paper a lean algorithm that is inspired by the bi-exponential filter and preserves its structure—a pair of one-tap recursions. By a careful but simple local adaptation of the filter weights to the data, we are able to design an edge-preserving smoother that has a very low memory and computational footprint while requiring a trivial coding effort. We demonstrate that our filter (a bi-exponential edge-preserving smoother, or BEEPS) has formal links with the traditional bilateral filter. On a practical side, we observe that the BEEPS also produces images that are similar to those that would result from the bilateral filter, but at a much-reduced computational cost. The cost per pixel is constant and depends neither on the data nor on the filter parameters, not even on the degree of smoothing.
H. Kirshner, T. Pengo, N. Olivier, D. Sage, S. Manley, M. Unser,
A PSF-Based Approach to Biplane Calibration in 3D Super-Resolution Microscopy,
IEEE International Symposium on Biomedical Imaging (ISBI'12), Barcelona, Spain, May 2-5, 2012.
Super-resolution localization microscopy methods such as PALM and STORM have been shown to provide imaging with resolutions up to a few tens of nanometers while using relatively simple setups. Biplane PALM has extended the PALM technique to three-dimensions, by simultaneously using two imaging planes, with different focal depths. A key aspect in achieving good axial localization results is the alignment of the two planes. Currently available approaches assume that misaligned planes only result in scaling and rotation of the PSF pattern. We show in this work that this does not necessarily hold true, especially in the presence of refractive index mismatch between the different optical layers. Instead, we suggest a calibration algorithm that relies on a realistic PSF model and finds the affine transform that relates the two planes with respect to a point source in the object domain. Our calibration algorithm also determines the defocus distance between the planes.
D. Sage,
Analyse d'image scientifique, le monde ImageJ,
Flash informatique 2012.
ImageJ est un logiciel Java de traitement d'images et d'analyse d'images scientifiques qui est dans le domaine public. ImageJ a un impact important en imagerie biomédicale, devenant de facto une référence. Son architecture logicielle ouverte a permis la réalisation de centaines de plugins intégrant parfois les techniques de traitement d'images les plus avancées.
R. Rezakhaniha, A. Agianniotis, J.T.C. Schrauwen, A. Griffa, D. Sage, C.V.C. Bouten, F.N. van de Vosse, M. Unser, N. Stergiopulos,
Experimental Investigation of Collagen Waviness and Orientation in the Arterial Adventitia Using Confocal Laser Scanning Microscopy,
Biomechanics and Modeling in Mechanobiology 11, 2012.
Mechanical properties of the adventitia are largely determined by the organization of collagen fibers. Measurements on the waviness and orientation of collagen, particularly at the zero-stress state, are necessary to relate the structural organization of collagen to the mechanical response of the adventitia. Using the fluorescence collagen marker CNA38-OG488 and confocal laser scanning microscopy, we imaged collagen fibers in the adventitia of rabbit common carotid arteries ex vivo. The arteries were cut open along their longitudinal axes to get the zero-stress state. We used semi-manual and automatic techniques to measure parameters related to the waviness and orientation of fibers. Our results showed that the straightness parameter (defined as the ratio between the distances of endpoints of a fiber to its length) was distributed with a beta distribution (mean value 0.72, variance 0.028) and did not depend on the mean angle orientation of fibers. Local angular density distributions revealed four axially symmetric families of fibers with mean directions of 0°, 90°, 43° and −43°, with respect to the axial direction of the artery, and corresponding circular standard deviations of 40°, 47°, 37° and 37°. The distribution of local orientations was shifted to the circumferential direction when measured in arteries at the zero-load state (intact), as compared to arteries at the zero-stress state (cut open). Information on collagen fiber waviness and orientation, such as obtained in this study, could be used to develop structural models of the adventitia, providing better means for analyzing and understanding the mechanical properties of vascular wall.
K.N. Chaudhury, D. Sage, M. Unser,
Fast O(1) Bilateral Filtering Using Trigonometric Range Kernels,
IEEE Transactions on Image Processing 20, 2011.
It is well known that spatial averaging can be realized (in space or frequency domain) using algorithms whose complexity does not scale with the size or shape of the filter. These fast algorithms are generally referred to as constant-time or O(1) algorithms in the image-processing literature. Along with the spatial filter, the edge-preserving bilateral filter involves an additional range kernel. This is used to restrict the averaging to those neighborhood pixels whose intensity are similar or close to that of the pixel of interest. The range kernel operates by acting on the pixel intensities. This makes the averaging process nonlinear and computationally intensive, particularly when the spatial filter is large. In this paper, we show how the O(1) averaging algorithms can be leveraged for realizing the bilateral filter in constant time, by using trigonometric range kernels. This is done by generalizing the idea presented by Porikli, i.e., using polynomial kernels. The class of trigonometric kernels turns out to be sufficiently rich, allowing for the approximation of the standard Gaussian bilateral filter. The attractive feature of our approach is that, for a fixed number of terms, the quality of approximation achieved using trigonometric kernels is much superior to that obtained by Porikli using polynomials.
D. Wüstner, J.R. Brewer, L. Bagatolli, D. Sage,
Potential of Ultraviolet Wide-Field Imaging and Multiphoton Microscopy for Analysis of Dehydroergosterol in Cellular Membranes,
Microscopy Research and Technique 74, 2011.
Dehydroergosterol (DHE) is an intrinsically fluorescent sterol with absorption/emission in the ultraviolet (UV) region and biophysical properties similar to those of cholesterol. We compared the potential of UV-sensitive low-light-level wide-field (UV-WF) imaging with that of multiphoton (MP) excitation microscopy to monitor DHE in living cells. Significantly reduced photobleaching in MP microscopy of DHE enabled us to acquire three-dimensional z-stacks of DHE-stained cells and to obtain high-resolution maps of DHE in surface ruffles, nanotubes, and the apical membrane of epithelial cells. We found that the lateral resolution of MP microscopy is ∼1.5-fold higher than that of UV-WF deconvolution microscopy, allowing for improved spatiotemporal analysis of plasma membrane sterol distribution. Surface intensity patterns of DHE with a diameter of 0.2 μm persisting over several minutes could be resolved by MP time-lapse microscopy. Diffusion coefficients of 0.25-μm-diameter endocytic vesicles containing DHE were determined by MP spatiotemporal image correlation spectroscopy. The requirement of extremely high laser power for visualization of DHE by MP microscopy made this method less potent for multicolor applications with organelle markers like green fluorescent protein-tagged proteins. The signal-to-noise ratio obtainable by UV-WF imaging could be significantly improved by pixelwise bleach rate fitting and calculation of an amplitude image from the decay model and by frame averaging after pixelwise bleaching correction of the image stacks. We conclude that UV-WF imaging and MP microscopy of DHE provide complementary information regarding membrane distribution and intracellular targeting of sterols.
D. Jeanbourquin, D. Sage, L. Nguyen, B. Schaeli, S. Kayal, D.A. Barry, L. Rossi,
Flow Measurements in Sewers Based on Image Analysis: Automatic Flow Velocity Algorithm,
Water Science & Technology 64, 2011.
Discharges of combined sewer overflows (CSOs) and stormwater are recognized as an important source of environmental contamination. However, the harsh sewer environment and particular hydraulic conditions during rain events reduce the reliability of traditional flow measurement probes. An in situ system for sewer water flow monitoring based on video images was evaluated. Algorithms to determine water velocities were developed based on image-processing techniques. The image-based water velocity algorithm identifies surface features and measures their positions with respect to real world coordinates. A web-based user interface and a three-tier system architecture enable remote configuration of the cameras and the image-processing algorithms in order to calculate automatically flow velocity on-line. Results of investigations conducted in a CSO are presented. The system was found to measure reliably water velocities, thereby providing the means to understand particular hydraulic behaviors.
A. Griffa, N. Garin, D. Sage,
Comparison of Deconvolution Software: A User Point of View—Part 2,
G.I.T. Imaging & Microscopy 12, 2010.
Deconvolution is an image processing technique that restores the effective object representation [3] [4], allowing to improve images analysis steps such as segmentation [1] or colocalization study [2]. We performed several deconvolution tests on different kinds of datasets. The methodology has been reported in Part 1. Evaluation criteria and results are reported here. References
A.F. Stalder, T. Melchior, M. Müller, D. Sage, T. Blu, M. Unser,
Low-Bond Axisymmetric Drop Shape Analysis for Surface Tension and Contact Angle Measurements of Sessile Drops,
Colloids and Surfaces A, 2010.
A new method based on the Young-Laplace equation for measuring contact angles and surface tensions is presented. In this approach, a first-order perturbation technique helps to analytically solve the Young-Laplace equation according to photographic images of axisymmetric sessile drops. When appropriate, the calculated drop contour is extended by mirror symmetry so that reflection of the drop into substrate allows the detection of position of the contact points. To keep a wide range of applicability, a discretisation of the drop's profile is not realised; instead, an optimisation of an advanced image-energy term fits an approximation of the Young-Laplace equation to drop boundaries. In addition, cubic B-spline interpolation is applied to the image of the drop to reach subpixel resolution. To demonstrate the method's accuracy, simulated drops as well as images of liquid coal ash slags were analysed. Thanks to the high-quality image interpolation model and the image-energy term, the experiments demonstrated robust measurements over a wide variety of image types and qualities. The method was implemented in Java and is freely available.
D. Sage, M. Unser, P. Salmon, C. Dibner,
A Software Solution for Recording Circadian Oscillator Features in Time-Lapse Live Cell Microscopy,
Cell Division 5, 2010.
Background: Fluorescent and bioluminescent time-lapse microscopy approaches have been successfully used to investigate molecular mechanisms underlying the mammalian circadian oscillator at the single cell level. However, most of the available software and common methods based on intensity-threshold segmentation and frame-to-frame tracking are not applicable in these experiments. This is due to cell movement and dramatic changes in the fluorescent/bioluminescent reporter protein during the circadian cycle, with the lowest expression level very close to the background intensity. At present, the standard approach to analyze data sets obtained from time lapse microscopy is either manual tracking or application of generic image-processing software/ dedicated tracking software. The latter however only applicable in cases where the cell does not change its position during the whole experiment, and thus are limiting. Results: In an attempt to improve existing methodology of time-lapse tracking of a large number of moving cells, we have developed a semi-automatic software package. It extracts the trajectory of the cells by tracking theirs displacements, makes the delineation of cell nucleus or whole cell, and finally yields measurements of various features, like reporter protein expression level or cell displacement. As an example, we present here single cell circadian pattern and motility analysis of NIH3T3 mouse fibroblasts expressing a fluorescent circadian reporter protein. Using Circadian Gene Express plugin, we performed fast and nonbiased analysis of large fluorescent time lapse microscopy datasets. Conclusions: Our software solution, Circadian Gene Express (CGE), is easy to use and allows precise and semi-automatic tracking of moving cells over longer period of time. In spite of significant circadian variations in protein expression with extremely low expression levels at the valley phase, CGE allows accurate and efficient recording of large number of cell parameters, including level of reporter protein expression, velocity, direction of movement, and others. CGE proves to be useful for the analysis of widefield fluorescent microscopy datasets, as well as for bioluminescence imaging. Moreover, it might be easily adaptable for confocal image analysis by manually choosing one of the focal planes of each z-stack of the various time points of a time series.
D. Wüstner, D. Sage,
Multicolor Bleach-Rate Imaging Enlightens in vivo Sterol Transport,
Communicative & Integrative Biology 3, 2010.
Elucidation of in vivo cholesterol transport and its aberrations in cardiovascular diseases requires suitable model organisms and the development of appropriate monitoring technology. We recently presented a new approach to visualize transport of the intrinsically fluorescent sterol, dehydroergosterol (DHE) in the genetically tractable model organism Caenorhabditis elegans (C. elegans). DHE is structurally very similar to cholesterol and ergosterol, two sterols used by the sterol-auxotroph nematode. We developed a new computational method measuring fluorophore bleaching kinetics at every pixel position, which can be used as a fingerprint to distinguish rapidly bleaching DHE from slowly bleaching autofluorescence in the animals. Here, we introduce multicolor bleach-rate sterol imaging. By this method, we demonstrate that some DHE is targeted to a population of basolateral recycling endosomes (RE) labelled with GFP-tagged RME-1 (GFP-RME-1) in the intestine of both, wild-type nematodes and mutant animals lacking intestinal gut granules (glo1-mutants). DHE-enriched intestinal organelles of glo1-mutants were decorated with GFP-rme8, a marker for early endosomes. No co-localization was found with a lysosomal marker, GFP-LMP1. Our new methods hold great promise for further studies on endosomal sterol transport in C. elegans.
D. Jeanbourquin, D. Sage, L. Nguyen, B. Schaeli, S. Kayal, D.A. Barry, L. Rossi,
Flow Measurements in Sewer Systems Based on Image Analysis: Automatic Flow Velocity Algorithm,
Int. Conf. on Sustainable Techniques and Strategies in Urban Water Management (NOVATECH), Lyon, France, 2010.
Discharges of combined sewer overflows (CSOs) and stormwater are recognized as an important source of environmental contamination. However, the harsh sewer environment and particular hydraulic conditions during rain events reduce the reliability of traditional flow measurement probes. An in situ system for sewer water flow monitoring based on video images was evaluated. Algorithm to determine water velocities was developed based on image-processing techniques. The image-based water velocity algorithm identifies surface features and measures their positions with respect to real world coordinates. A web-based user interface and a three-tier system architecture enable remote configuration of the cameras and the image-processing algorithms in order to calculate automatically flow velocity on-line. Results of investigations conducted in a CSO were presented. The system was found to measure reliably water velocities, thereby providing the means to understand particular hydraulic behaviors.
K.N. Chaudhury, Z. Püspöki, A. Muñoz Barrutia, D. Sage, M. Unser,
Fast Detection of Cells Using a Continuously Scalable Mexican-Hat-Like Template,
IEEE International Symposium on Biomedical Imaging (ISBI), 2010.
We propose a fast algorithm for the detection of cells in fluorescence images. The algorithm, which estimates the number of cells and their respective centers and radii, relies on the fast computation of intensity-based correlations between the cells and a near-isotropic Mexican-hat-like detector. The attractive features of our algorithm are its speed and accuracy. The former attribute is derived from the fact that we can compute correlations between a cell and detectors of various sizes using O(1) operations; whereas, it is our ability to continuously control the center and the radius of the detector that results in a precise estimate of the position and size of the cell. We provide experimental results on both simulated and real data to demonstrate the speed and accuracy of the algorithm.
D. Wüstner, A.L. Larsen, N.J. Færgeman, J.R. Brewer, D. Sage,
Selective Visualization of Fluorescent Sterols in Caenorhabditis elegans by Bleach-Rate-Based Image Segmentation,
Traffic 11, 2010.
The nematode Caenorhabditis elegans is a genetically tractable model organism to investigate sterol transport. In vivo imaging of the fluorescent sterol, dehydroergosterol (DHE), is challenged by C. elegans' high autofluorescence in the same spectral region as emission of DHE. We present a method to detect DHE selectively, based on its rapid bleaching kinetics compared to cellular autofluorescence. Worms were repeatedly imaged on an ultraviolet-sensitive wide field (UV-WF) microscope, and bleaching kinetics of DHE were fitted on a pixel-basis to mathematical models describing the intensity decay. Bleach-rate constants were determined for DHE in vivo and confirmed in model membranes. Using this method, we could detect enrichment of DHE in specific tissues like the nerve ring, the spermateca and oocytes. We confirm these results in C. elegans gut-granule-loss (glo) mutants with reduced autofluorescence and compare our method with three-photon excitation microscopy of sterol in selected tissues. Bleach-rate-based UV-WF imaging is a useful tool for genetic screening experiments on sterol transport, as exemplified by RNA interference against the rme-2 gene coding for the yolk receptor and for worm homologues of Niemann-Pick C disease proteins. Our approach is generally useful for identifying fluorescent probes in the presence of high cellular autofluorescence.
A. Griffa, N. Garin, D. Sage,
Comparison of Deconvolution Software in 3D Microscopy: A User Point of View—Part 1,
G.I.T. Imaging & Microscopy 12, 2010.
Deconvolution is an image restoration technique which improves image contrast, resolution and signal to noise ratio. In modern optical microscopy and biological research deconvolution is becoming a fundamental processing step which allows for better image analysis. Deconvolution remains however a challenging task as the result depends strongly on the algorithm chosen, the parameters settings and the kinds of structures in the processed dataset. As a core facility of bio-imaging and microscopy, we aim with this study to compare the performances of different deconvolution software. In this first part of our survey we present deconvolution related problems, we introduce software we took into account, and we provide the complete dataset we produced for software testing and a PSF generator. A second part will follow the present one. In the second part we will highlight advantages and weak points of tested software by the statement of the performed tests.
M. Unser, D. Sage, D. Van De Ville,
Multiresolution Monogenic Signal Analysis Using the Riesz-Laplace Wavelet Transform,
IEEE Transactions on Image Processing 18, 2009.
The monogenic signal is the natural 2-D counterpart of the 1-D analytic signal. We propose to transpose the concept to the wavelet domain by considering a complexified version of the Riesz transform which has the remarkable property of mapping a real-valued (primary) wavelet basis of L2(ℝ2) into a complex one. The Riesz operator is also steerable in the sense that it give access to the Hilbert transform of the signal along any orientation. Having set those foundations, we specify a primary polyharmonic spline wavelet basis of L2(ℝ2) that involves a single Mexican-hat-like mother wavelet (Laplacian of a B-spline). The important point is that our primary wavelets are quasi-isotropic: they behave like multiscale versions of the fractional Laplace operator from which they are derived, which ensures steerability. We propose to pair these real-valued basis functions with their complex Riesz counterparts to specify a multiresolution monogenic signal analysis. This yields a representation where each wavelet index is associated with a local orientation, an amplitude and a phase. We give a corresponding wavelet-domain method for estimating the underlying instantaneous frequency. We also provide a mechanism for improving the shift and rotation-invariance of the wavelet decomposition and show how to implement the transform efficiently using perfect-reconstruction filterbanks. We illustrate the specific feature-extraction capabilities of the representation and present novel examples of wavelet-domain processing; in particular, a robust, tensor-based analysis of directional image patterns, the demodulation of interferograms, and the reconstruction of digital holograms.
E. Fonck, G.G. Feigl, J. Fasel, D. Sage, M. Unser, D.A. Rüfenacht, N. Stergiopulos,
Effect of Aging on Elastin Functionality in Human Cerebral Arteries,
Stroke 40, 2009.
Background and Purpose—Aging affects elastin, a key component of the arterial wall integrity and functionality. Elastin degradation in cerebral vessels is associated with cerebrovascular disease. The goal of this study is to assess the biomechanical properties of human cerebral arteries, their composition, and their geometry, with particular focus on the functional alteration of elastin attributable to aging. Methods—Twelve posterior cranial arteries obtained from human cadavers of 2 different age groups were compared morphologically and tested biomechanically before and after enzymatic degradation of elastin. Light, confocal, and scanning electron microscopy were used to analyze and determine structural differences, potentially attributed to aging. Results—Aging affects structural morphology and the mechanical properties of intracranial arteries. In contrast to main systemic arteries, intima and media thicken while outer diameter remains relatively constant with age, leading to concentric hypertrophy. The structural morphology of elastin changed from a fiber network oriented primarily in the circumferential direction to a more heterogeneously oriented fiber mesh, especially at the intima. Biomechanically, cerebral arteries stiffen with age and lose compliance in the elastin dominated regime. Enzymatic degradation of elastin led to loss in compliance and stiffening in the young group but did not affect the structural and material properties in the older group, suggesting that elastin, though present in equal quantities in the old group, becomes dysfunctional with aging. Conclusions—Elastin loses its functionality in cerebral arteries with aging, leading to stiffer less compliant arteries. The area fraction of elastin remained, however, fairly constant. The loss of functionality may thus be attributed to fragmentation and structural reorganization of elastin occurring with age.
L. Nguyen, D. Sage, S. Kayal, D. Jeanbourquin, M. Scherz, D.A. Barry, M. Unser, L. Rossi,
Vision-Based System for the Control and Measurement of Wastewater Flow Rate in Sewer Systems,
IWA Conference on Instrumentation, Control and Automation (ICA), 2009.
Total pollutant mass carried by sewer systems is an important factor affecting the quality of receiving waters [Burton and Pitt 2002]. This is especially the case during sizable rain events, when the total amount of the wastewater/stormwater mixture cannot be treated in the wastewater treatment plant; the overflow water goes directly into the environment (streams, rivers, lakes, coastal seas) through combined sewer overflows (CSOs). This water is untreated and is recognized as an important source of pollution [e.g., Even et al. 2006, Chèvre et al. 2007, Rossi et al. 2009]. Reliable pollutant concentration data are often available via sampling and analysis. However, accurate estimation of the total pollutant mass is infeasible unless the total volumetric flow rate is known with precision and reliability, and so the environmental impact due to CSO's cannot be predicted. Knowledge of volumetric water flow rate in sewer systems is a key environmental parameter. Robust monitoring of sewers is a challenging task as the environment within them is inherently harsh: constant humidity of 100%, rapid and large water level changes, corrosive atmosphere, presence of gas, difficult access and solid debris inside the flow are some of the usual problems. Accurate monitoring of the volumetric water flow is thus of crucial environmental importance. In this paper, a novel approach to the monitoring of flow in sewers based on video images is presented. Specifically, the method is based on image processing and computer vision techniques for water level and flow velocity estimation. References
.A. da Silveira, B.L. Schneider, C. Cifuentes-Diaz, D. Sage, T. Abbas-Terki, T. Iwatsubo, M. Unser, P. Aebischer,
Phosphorylation Does Not Prompt, Nor Prevent, the Formation of α-Synuclein Toxic Species in a Rat Model of Parkinson's Disease,
Human Molecular Genetics 18, 2009.
Phosphorylation is involved in numerous neurodegenerative diseases. In particular, alpha-synuclein is extensively phosphorylated in aggregates in patients suffering from synucleinopathies. However, the share of this modification in the events that lead to the conversion of alpha-synuclein to aggregated toxic species needed to be clarified. The rat model that we developed through rAAV2/6-mediated expression of alpha-synuclein demonstrates a correlation between neurodegeneration and formation of small filamentous alpha-synuclein aggregates. A mutation preventing phosphorylation (S129A) significantly increases alphasynuclein toxicity and leads to enhanced formation of beta-sheet-rich, proteinase K-resistant aggregates, increased affinity for intracellular membranes, a disarrayed network of neurofilaments and enhanced alpha-synuclein nuclear localization. The expression of a mutation mimicking phosphorylation (S129D) does not lead to dopaminergic cell loss. Nevertheless, fewer but larger aggregates are formed, and signals of apoptosis are also activated in rats expressing the phosphorylation-mimicking form of alpha-synuclein. These observations strongly suggest that phosphorylation does not play an active role in the accumulation of cytotoxic pre-inclusion aggregates. Unexpectedly, the study also demonstrates that constitutive expression of phosphorylation-mimicking forms of alpha-synuclein does not protect from neurodegeneration. The role of phosphorylation at Serine 129 in the early phase of Parkinson's disease is examined, which brings new perspective to therapeutic approaches focusing on the modulation of kinases/phosphatases activity to control alpha-synuclein toxicity.
C. Dibner, D. Sage, M. Unser, C. Bauer, T. d'Eysmond, F. Naef, Ü. Schibler,
Circadian Gene Expression Is Resilient to Large Fluctuations in Overall Transcription Rates,
The European Molecular Biology Organization Journal 28, 2009.
Mammalian circadian oscillators are considered to rely on transcription/translation feedback loops in clock gene expression. The major and essential loop involves the autorepression of cryptochrome (Cry1, Cry2) and period (Per1, Per2) genes. The rhythm-generating circuitry is functional in most cell types, including cultured fibroblasts. Using this system, we show that significant reduction in RNA polymerase II-dependent transcription did not abolish circadian oscillations, but surprisingly accelerated them. A similar period shortening was observed at reduced incubation temperatures in wild-type mouse fibroblasts, but not in cells lacking Per1. Our data suggest that mammalian circadian oscillators are resilient to large fluctuations in general transcription rates and temperature, and that PER1 has an important function in transcription and temperature compensation.
L.S. Nguyen, B. Schaeli, D. Sage, S. Kayal, D. Jeanbourquin, D.A. Barry, L. Rossi,
Vision-Based System for the Control and Measurement of Wastewater Flow Rate in Sewer Systems,
Water Science & Technology 60, 2009.
Combined sewer overflows and stormwater discharges represent an important source of contamination to the environment. However, the harsh environment inside sewers and particular hydraulic conditions during rain events reduce the reliability of traditional flow measurement probes. In the following, we present and evaluate an in situ system for the monitoring of water flow in sewers based on video images. This paper focuses on the measurement of the water level based on image-processing techniques. The developed image-based water level algorithms identify the wall/water interface from sewer images and measure its position with respect to real world coordinates. A web-based user interface and a 3-tier system architecture enable the remote configuration of the cameras and the image-processing algorithms. Images acquired and processed by our system were found to reliably measure water levels and thereby to provide crucial information leading to better understand particular hydraulic behaviors. In terms of robustness and accuracy, the water level algorithm provided equal or better results compared to traditional water level probes in three different in situ configurations.
D. Van De Ville, D. Sage, K. Balać, M. Unser,
The Marr Wavelet Pyramid and Multiscale Directional Image Analysis,
European Signal Processing Conference (EUSIPCO), 2008.
The “Marr wavelet pyramid” is a wavelet decomposition that implements a multiscale version of the complex gradient-Laplace operator. It is closely linked to a multiresolution analysis of L2(ℝ2) and it has a fast filterbank implementation. We show how the Marr wavelets, which are essentially steerable, can be used to extract a multiscale version of the structure tensor. This yields a multiscale characterization of an image in terms of various features such as local gradient energy, orientation, and coherency. We provide an implementation of the proposed system as a Java plug-in for ImageJ, and we illustrate its applicability to directional image analysis which is useful in domains such as biological imaging and material science.
E. Fonck, G.G. Feigl, J. Fasel, D. Sage, M. Unser, D.A. Rüfenacht, N. Stergiopulos,
Effect of Ageing on Elastin Functionality in Human Cerebral Arteries,
ASME 2008 Summer Bioengineering Conference (SBC), Marco Island, USA, 2008.
The ageing process affects elastin, a key component of the arterial wall integrity and functionality. Elastin may play an important role in cerebral vessels because elastin degradation is linked to cerebrovascular disease [1]. The goal of this study is to assess the biomechanical properties of human cerebral arteries, their composition and geometry, with particular focus on the functional alterations of elastin in cerebral arteries due to ageing.
M. Tsukahara, S. Mitrović, V. Gajdosik, G. Margaritondo, L. Pournin, M. Ramaioli, D. Sage, Y. Hwu, M. Unser, T.M. Liebling,
Coupled Tomography and Distinct-Element-Method Approach to Exploring the Granular Media Microstructure in a Jamming Hourglass,
Physical Review E 77, 2008.
We describe an approach for exploring microscopic properties of granular media that couples x-ray microtomography and distinct-element-method (DEM) simulations through image analysis. We illustrate it via the study of the intriguing phenomenon of instant arching in an hourglass (in our case a cylinder filled with a polydisperse mixture of glass beads that has a small circular shutter in the bottom). X-ray tomography provides three-dimensional snapshots of the microscopic conditions of the system both prior to opening the shutter, and thereafter, once jamming is completed. The process time in between is bridged using DEM simulation, which settles to positions in remarkably good agreement with the x-ray images. Specifically designed image analysis procedures accurately extract the geometrical information, i.e., the positions and sizes of the beads, from the raw x-ray tomographs, and compress the data representation from initially 5 gigabytes to a few tens of kilobytes per tomograph. The scope of the approach is explored through a sensitivity analysis to input data perturbations in both bead sizes and positions. We establish that accuracy of size—much more than position—estimates is critical, thus explaining the difficulty in considering a mixture of beads of different sizes. We further point to limits in the replication ability of granular flows away from equilibrium; i.e., the difficulty of numerically reproducing chaotic motion.
H. Schober, V. Kalck, M.A. Vega-Palas, G. Van Houwe, D. Sage, M. Unser, M.R. Gartenberg, S.M. Gasser,
Controlled Exchange of Chromosomal Arms Reveals Principles Driving Telomere Interactions in Yeast,
Genome Research 18, 2008.
The 32 telomeres in the budding yeast genome cluster in three to seven perinuclear foci. Although individual telomeres and telomeric foci are in constant motion, preferential juxtaposition of some telomeres has been scored. To examine the principles that guide such long-range interactions, we differentially tagged pairs of chromosome ends and developed an automated three-dimensional measuring tool that determines distances between two telomeres. In yeast, all chromosomal ends terminate in TG1-3 and middle repetitive elements, yet subgroups of telomeres also share extensive homology in subtelomeric coding domains. We find that up to 21 kb of >90% sequence identity does not promote telomere pairing in interphase cells. To test whether unique sequence elements, arm length, or chromosome territories influence juxtaposition, we reciprocally swapped terminal domains or entire chromosomal arms from one chromosome to another. We find that the distal 10 kb of Tel6R promotes interaction with Tel6L, yet only when the two telomeres are present on the same chromosome. By manipulating the length and sequence composition of the right arm of chr 5, we confirm that contact between telomeres on opposite chromatid arms of equal length is favored. These results can be explained by the polarized Rabl arrangement of yeast centromeres and telomeres, which promote to telomere pairing by allowing contact between chromosome arms of equal length in anaphase.
A. Depeursinge, D. Sage, A. Hidki, A. Platon, P.-A. Poletti, M. Unser, H. Müller,
Lung Tissue Classification Using Wavelet Frames,
Int. Conf. of the IEEE Engineering in Medicine and Biology Society, 2007.
We describe a texture classification system that identifies lung tissue patterns from high-resolution computed tomography (HRCT) images of patients affected with interstitial lung diseases (ILD). This pattern recognition task is part of an image-based diagnostic aid system for ILDs. Five lung tissue patterns (healthy, emphysema, ground glass, fibrosis and microdules) selected from a multimedia database are classified using the overcomplete discrete wavelet frame decompostion combined with grey-level histogram features. The overall multiclass accuracy reaches 92.5% of correct matches while combining the two types of features, which are found to be complementary.
A.F. Stalder, G. Kulik, D. Sage, L. Barbieri, P. Hoffmann,
A Snake-Based Approach to Accurate Determination of Both Contact Points and Contact Angles,
Colloids And Surfaces A 286, 2006.
We present a new method based on B-spline snakes (active contours) for measuring high-accuracy contact angles. In this approach, we avoid making physical assumptions by defining the contour of the drop as a versatile B-spline curve. When useful, we extend this curve by mirror symmetry so that we can take advantage of the reflection of the drop onto the substrate to detect the position of the contact points. To keep a wide range of applicability, we refrain from discretizing the contour of the drop, and we choose to optimize an advanced image-energy term to drive the evolution of the curve. This term has directional gradient and region-based components; additionally, another term—an internal energy—is responsible for the snake elasticity and constrains the parameterization of the spline. While preserving precision at the contact points, we limit the computational complexity by constraining a non-uniform repartition of the control points. The elasticity property of the snake links the local nature of the contact angle to the global contour of the drop. A global knowledge of the drop contour allows us to use the reflection of the drop on the substrate to automatically and precisely detect a line of contact points (vertical position and tilt). We apply cubic-spline interpolation over the image of the drop; then, the evolution procedure takes part in this continuous domain to avoid the inaccuracies introduced by pixelization and discretization. We have programmed our method as a Java software and we make it freely available. Our experiments result in good accuracy thanks to our high-quality image-interpolation model, while they show applicability to a variety of images thanks to our advanced image-energy term.
A. Hirabayashi, D. Sage, M. Unser,
IP-LAB: A Tool to Teach Image Processing Programming in Java—Challenge in Swiss Federal Institute of Technology Lausanne,
Fifty-Annual Japanese Society for Engineering Education Conference (JSEE), 2005.
Image processing is getting more and more important because of the widespread use of digital imaging devices, such as digital cameras or digital videos. While image processing is a very practical discipline, it is perceived as being rather theoretical. It is actually a subject that involves a rigorous, mathematical treatment. Hence, what is important in image processing education is that instructors make students not only understand such mathematical formulas but also become to be able to use them. To this end, computer laboratories accompanied by lectures are very effective for image processing education. In order to make the best use of them, computer systems which will be used by students play a very important role. IP-Lab is a system in Java that was developed by the second and the third authors of the present paper in the Swiss Federal Institute of Technology Lausanne (EPFL) to offer an environment where the students could implement the algorithms literally as they are seen in the course.
J.N. Feige, D. Sage, W. Wahli, B. Desvergne, L. Gelman,
PixFRET, an ImageJ Plug-in for FRET Calculation That Can Accommodate Variations in Spectral Bleed-throughs,
Microscopy Research and Technique 68, 2005.
Fluorescence resonance energy transfer (FRET) allows the user to investigate interactions between fluorescent partners. One crucial issue when calculating sensitized emission FRET is the correction for spectral bleed-throughs (SBTs), which requires to calculate the ratios between the intensities in the FRET and in the donor or acceptor settings, when only the donor or acceptor are present. Theoretically, SBT ratios should be constant. However, experimentally, these ratios can vary as a function of fluorophore intensity, and assuming constant values may hinder precise FRET calculation. One possible cause for such a variation is the use of a microscope set-up with different photomultipliers for the donor and FRET channels, a set-up allowing higher speed acquisitions on very dynamic fluorescent molecules in living cells. Herein, we show that the bias introduced by the differential response of the two PMTs can be circumvented by a simple modeling of the SBT ratios as a function of fluorophore intensity. Another important issue when performing FRET is the localization of FRET within the cell or a population of cells. We hence developed a freely available ImageJ plug-in, called PixFRET, that allows a simple and rapid determination of SBT parameters and the display of normalized FRET images. The usefulness of this modeling and of the plug-in are exemplified by the study of FRET in a system where two interacting nuclear receptors labeled with ECFP and EYFP are coexpressed in living cells.
D. Sage, F.R. Neumann, F. Hediger, S.M. Gasser, M. Unser,
Automatic Tracking of Individual Fluorescence Particles: Application to the Study of Chromosome Dynamics,
IEEE Transactions on Image Processing 14, 2005.
We present a new, robust, computational procedure for tracking fluorescent markers in time-lapse microscopy. The algorithm is optimized for finding the time-trajectory of single particles in very noisy dynamic (two- or three-dimensional) image sequences. It proceeds in three steps. First, the images are aligned to compensate for the movement of the biological structure under investigation. Second, the particle's signature is enhanced by applying a Mexican hat filter, which we show to be the optimal detector of a Gaussian-like spot in 1 ⁄ ω2 noise. Finally, the optimal trajectory of the particle is extracted by applying a dynamic programming optimization procedure. We have used this software, which is implemented as a Java plug-in for the public-domain ImageJ software, to track the movement of chromosomal loci within nuclei of budding yeast cells. Besides reducing trajectory analysis time by several 100-fold, we achieve high reproducibility and accuracy of tracking. The application of the method to yeast chromatin dynamics reveals different classes of constraints on mobility of telomeres, reflecting differences in nuclear envelope association. The generic nature of the software allows application to a variety of similar biological imaging tasks that require the extraction and quantitation of a moving particle's trajectory.
U. Rajashekar, A.C. Bovik, D. Sage, M. Unser, L.J. Karam, R.L. Lagendijk,
Image Processing Education,
Handbook of Image & Video Processing A. Bovik, 2005.
This chapter describes tools and techniques that facilitate a gentle introduction to fascinating concepts in digital image processing from four leading universities. Equipped with informative visualizations and a user-friendly interfaces, these modules are currently being used effectively in a classroom environment for teaching DIP at many universities across the world.
B. Forster, D. Van De Ville, J. Berent, D. Sage, M. Unser,
Complex Wavelets for Extended Depth-of-Field: A New Method for the Fusion of Multichannel Microscopy Images,
Microscopy Research and Technique 65, 2004.
Microscopy imaging often suffers from limited depth-of-field. However, the specimen can be “optically sectioned” by moving the object along the optical axis. Then different areas appear in focus in different images. Extended depth-of-field is a fusion algorithm that combines those images into one single sharp composite. One promising method is based on the wavelet transform. Here, we show how the wavelet-based image fusion technique can be improved and easily extended to multichannel data. First, we propose the use of complex-valued wavelet bases, which seem to outperform traditional real-valued wavelet transforms. Second, we introduce a way to apply this technique for multichannel images that suppresses artifacts and does not introduce false colors, an important requirement for multichannel optical microscopy imaging. We evaluate our method on simulated image stacks and give results relevant to biological imaging.
B. Forster, D. Van De Ville, J. Berent, D. Sage, M. Unser,
Extended Depth-of-Focus for Multi-Channel Microscopy Images: A Complex Wavelet Approach,
IEEE International Symposium on Biomedical Imaging (ISBI), 2004.
Microscopy imaging often suffers from limited depth-of-focus. However, the specimen can be “optically sectioned” by moving the object along the optical axis; different areas appear in focus in different images. Extended depth-of-focus is a fusion algorithm that combines those images into one single sharp composite. One promising method is based on the wavelet transform. In this paper, we show how the wavelet-based image fusion technique can be improved and easily extended to multi-channel data. First, we propose the use of complex-valued wavelet bases, which seem to outperform traditional real-valued wavelet transforms. Second, we introduce a way to apply this technique for multi-channel images that suppresses artifacts and does not introduce false colors, an important requirement for multi-channel fluorescence microscopy imaging. We evaluate our method on simulated image stacks and give results relevant to biological imaging.
D. Sage, M. Unser,
Teaching Image-Processing Programming in Java,
IEEE Signal Processing Magazine 20, 2003.
Image processing (IP) can be taught very effectively by complementing the basic lectures with computer laboratories where the participants can actively manipulate and process images. This offering can be made even more attractive by allowing the students to develop their own IP code within a reasonable time frame. After a brief review of existing software packages that can be used for teaching IP, we present a system that we have designed to be as “student-friendly” as possible. The software is built around ImageJ, a freely available, full-featured, and user-friendly program for image analysis. The computer sessions are alternated with lectures, typically, a three-hour session at the end of every chapter. The sessions are in the form of assignments that guide the students towards the solution of simple imaging problems. The starting point is typically the understanding and testing of some standard IP algorithm in Java. Next, students are asked to extend the algorithms progressively. This constructive approach is made possible thanks to a programmer-friendly environment and an additional software interface layer that greatly facilitates the developments of plug-ins for ImageJ. Taking into account the fact that our students are not experienced programmers (they typically do not even know Java), we use a “learn by example” teaching strategy, with good success.
D. Sage, F. Hediger, S.M. Gasser, M. Unser,
Automatic Tracking of Particles in Dynamic Fluorescence Microscopy,
International Symposium on Image and Signal Processing and Analysis (ISPA), 2003.
We present a new, robust algorithm for tracking fluorescent particles in dynamic image sequences obtained by brightfield or confocal microscopy. Specifically, we consider the problem of extracting the movement of chromosomal telomeres within the nucleus of a budding yeast cell. Our method has three components. The first is an alignment module that compensates for the movement of the biological structure under investigation. In our application, the images are aligned to the center of gravity of the nucleus which is detected by thresholding and fitted with an ellipse. The second step is a Mexican-hat filtering which we show to be optimally tailored to the detection of a Gaussian-like spot in fractal noise. The final component is a tracking algorithm that uses dynamic programming to extract the optimal (x, y, t) trajectory of a particle. We have implemented the method as a Java Plugin for the public-domain ImageJ software. We have applied it to real data and have obtained results that are as good—if not better—as manual tracings. Our new algorithm reduces the analysis time of a 300 image sequence from 10 minutes, when it is done manually, to just a few seconds and offers the benefit of reproducibility.
D. Sage, M. Unser,
Easy Java Programming for Teaching Image Processing,
2001 IEEE International Conference on Image Processing (ICIP), 2001.
We have designed a series of computer sessions build around ImageJ (a public-domain software for image analysis), as a practical complement to a two-semester course in image processing. The students are challenged with simple practical imaging problems as they acquire hands-on practice by experimenting with image-processing operators. In the process, they also learn how to program standard image-processing algorithms in Java. This is made possible thanks to a programmer-friendly environment and a software interface that greatly facilitates the developments of plugins for ImageJ. Since our students have generally not acquired programming skills yet (they typically do not even know Java), we use a learning-by-example teaching strategy, with good success.